We asked the heads of our scientific divisions to tell us about some of the big questions in fundamental biomedical science that researchers are investigating with NIGMS support. This article is the second in an occasional series that explores these questions and explains how pursuing the answers could advance understanding of important biological processes.
For some health conditions, the cause is clear: A single altered gene is responsible. But for many others, the path to disease is more complex. Scientists are working to understand how factors like genetics, lifestyle and environmental exposures all contribute to disease. Another important, but less well-known, area of investigation is the role of chance at the molecular level.
One team working in this field is led by John Tyson at Virginia Tech. The group focuses on how chance events affect the cell division cycle, in which a cell duplicates its contents and splits into two. This cycle is the basis for normal growth, reproduction and the replenishment of skin, blood and other cells throughout the body. Errors in the cycle are associated with a number of conditions, including birth defects and cancer.
Tyson wants to understand how the cell cycle functions properly despite random fluctuations in the number of copies of one of its key players, messenger RNA (mRNA). mRNA molecules transmit information coded in genes to the cell’s protein factories. Cell cycle-related mRNA molecules enable the production of the proteins that make cell division possible.
Tyson first needed to figure out just how widely mRNA numbers vary. To do this, he collaborated with Virginia Tech computational biologist Jean Peccoud and his team at the Virginia Bioinformatics Institute. Together, they studied yeast cells, which use the same molecular machinery for cell division as human cells but are easier to work with because of their rapid growth and simple genome.
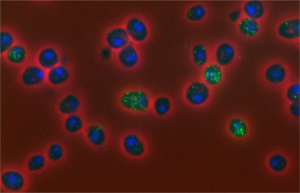
The researchers used a technique called fluorescence in situ hybridization (FISH) to count the number of mRNA copies in a cell. They placed a yeast cell on a microscope slide. They then exposed the cell to fluorescent molecules that stick to specific mRNAs. Next, like shaking glitter from a picture drawn in glue, they washed away the fluorescent molecules that didn’t stick and were left with glowing dots marking each mRNA. They did this for hundreds of yeast cells and found that most had around five or six copies of any given cell cycle-related mRNA molecule, but some had as few as one or as many as ten. A cell with ten copies of a particular mRNA molecule would be about 10 times more likely to turn the instructions encoded in that mRNA into a protein as a cell that only had one copy of the mRNA.
Tyson and Peccoud next wanted to explore the impact of mRNA variations on the yeast cell’s ability to grow and divide properly. Because this question is far too complex to answer with a single experiment, they created a mathematical model that could track how different numbers of mRNAs in yeast cells affect all of the many genes and proteins that make up the other important players in the cell cycle control machinery.
Their model predicts that when cell cycle-related mRNA levels fluctuate, the corresponding protein levels do as well. According to the model, the variation in protein number in healthy yeast cells is within a range that consistently allows the cell cycle to function reliably. With even slightly fewer mRNA molecules per cell on average, the machinery wouldn’t work as reliably in many cells.
Tyson and Peccoud now plan to explore in more detail how fluctuating mRNA levels affect one particular component of the cell cycle machinery called the spindle assembly checkpoint. This critical quality-control mechanism halts cell division until the duplicated DNA is neatly bundled into two matching sets of chromosomes ready to be pulled to opposite sides of the cell. If the spindle assembly checkpoint fails, divided cells can have too many or too few chromosomes, a condition often associated with cancer.
Tyson and Peccoud hope that their studies will improve understanding of whether and how chance events in cells contribute to both health and disease.


Could cells be duplicated and increased to outnumber those cells affected by viruses that are nearly impossible to combat?
Interesting idea! But that strategy probably wouldn’t work because most viruses replicate and spread faster than cells can divide.
Fascinating article, especially the section on spindle assembly checkpoints. It makes me wonder, what if there was a way to increase the resiliency of the checkpoints? I wonder if there’s any tie-in between angiogenesis towards tumors and failure of spindle assembly checkpoints. Not a SME by any stretch of the imagination, so if I made no sense, disregard.