Leafing through my favorite biology textbook from a handful of years ago, I was struck by the relative brevity of the chapter on human evolution. While other fields of biological research have enjoyed a steady gallop of productivity over the last few decades due in part to advances in computing power, imaging technology and experimental methods, the study of human evolution can be seen as having lagged behind until recently due to an almost complete dependence on fossil evidence.
Fortunately, contemporary biology textbook chapters on human evolution are being primed for a serious upgrade thanks to the recent availability of high-quality genome sequences from diverse modern human populations as well as from ancient humans and other non-human hominids, including the Neanderthals and Denisovans (but, for purposes of this story, not the Great Apes).
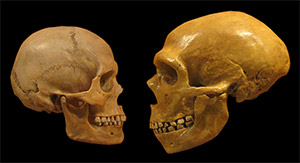
What are the new resources for studying human evolution?
The cost of DNA sequencing has dropped precipitously in the last decade. As a result, more complete human genome sequences become available for analysis with each passing year.
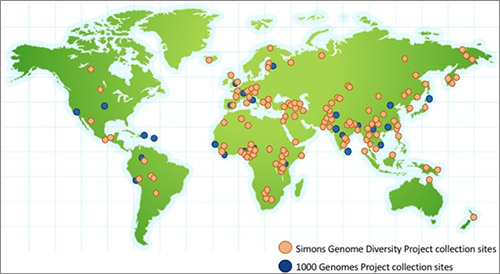
For example, the 1000 Genomes Project ![]() includes more than 1,000 full human genome sequences of individuals from European, Asian, American and Sub-Saharan African populations. Earlier this year, the Simons Genome Diversity Project
includes more than 1,000 full human genome sequences of individuals from European, Asian, American and Sub-Saharan African populations. Earlier this year, the Simons Genome Diversity Project ![]() further increased the number of available human genomes by adding 300 individuals representing 142 populations around the globe.
further increased the number of available human genomes by adding 300 individuals representing 142 populations around the globe.
Other high-quality genome sequences that scientists have recently made available include those of Aboriginal Australians as well as people from Papua, New Guinea, the Bismarck Islands and the Solomon Islands.
Ancient genomes have been sequenced from early human remains collected from archaeological sites in the South Pacific island nations of Vanuatu and Tonga, and other places.
Scientists have also sequenced the genomes of two ancient hominids, Denisovans and Neanderthals.
The availability of so many human and non-human hominid genomes with identified ancestry, as well as the advent of more powerful statistical methods for genome comparisons, have allowed researchers to ask deeper and more precise questions about the course of long-term human history. In addition, we continue to discover remains of additional ancient ancestors, such as Homo floresiensis in 2003 and Homo nadeli in 2013.
What has cataloging human genetic diversity revealed?
Studies of present and past genetic diversity address longstanding questions about the movement and diversification of human populations and the origins of culture.
The most dramatic human population genetic differences are among modern Africans. On the other hand, modern non-Africans can trace their ancestry back to a single population that moved out of Africa between 50,000 and 80,000 years ago and not, as described by a competing hypothesis, to several phases of migration dating back to 130,000 years ago. Check out the NIH Director’s Blog on this topic for more details.
Comparisons of modern and ancient genome sequences are giving a clearer picture of how humans are related to Neanderthals and Denisovans. These comparisons are also illuminating the history of humans interbreeding with non-human hominids. For example, more evidence of Neanderthal ancestry can be found in East Asians than in West Eurasians. This pattern may be the result of increased hybridization between Neanderthals and ancestral East Asians or by dilution of Neanderthal ancestry in ancestral West Eurasians due to their hybridization with other as yet unidentified populations.
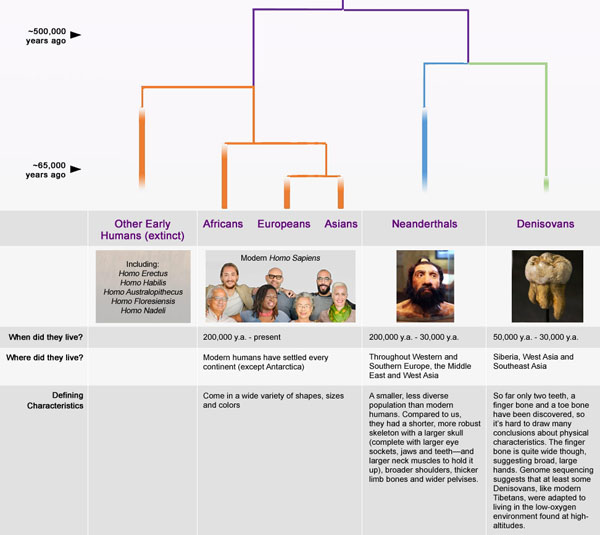
In addition, recent studies describe the timing of the separation of ancestral human populations that gave rise to modern populations. For example, the island cultures of the South Pacific represent the last significant migration of humans into lands that were previously uninhabited. Analyses of genome-wide ancient DNA from archaeological sites throughout the South Pacific are redrawing the likeliest route of migrants to South Pacific frontiers based on their sequence similarity to that of mainland East Asians, Papuans and other Pacific Islanders.
The timing of movement and separation of ancestral populations can also be linked to what is known about the adoption of culture. For example, the practice of farming, rather than scavenging or hunter-gathering, was introduced across the Near East at a faster rate than the spread of the population of known early farmers. In other words, the competency of farming spread faster than the early farmers did!
Studies such as these strike at the heart of what makes us human and how modern human diversity arose from a much smaller, more homogeneous population of ancestors less than 200,000 years ago.
More Biomedical Beat Stories



Great this is what I like. Human evolution.