As a medical student, Martin Burke, M.D., Ph.D., helped care for a young college student with cystic fibrosis (CF), an inherited disease that affects the body’s ability to make sweat and mucus. Dr. Burke had just studied CF in class, so he relayed what he had learned to her. He had a lot of information to give—doctors and researchers know the exact amino acid changes in an ion channel protein called cystic fibrosis transmembrane conductance regulator (CFTR) that cause CF.
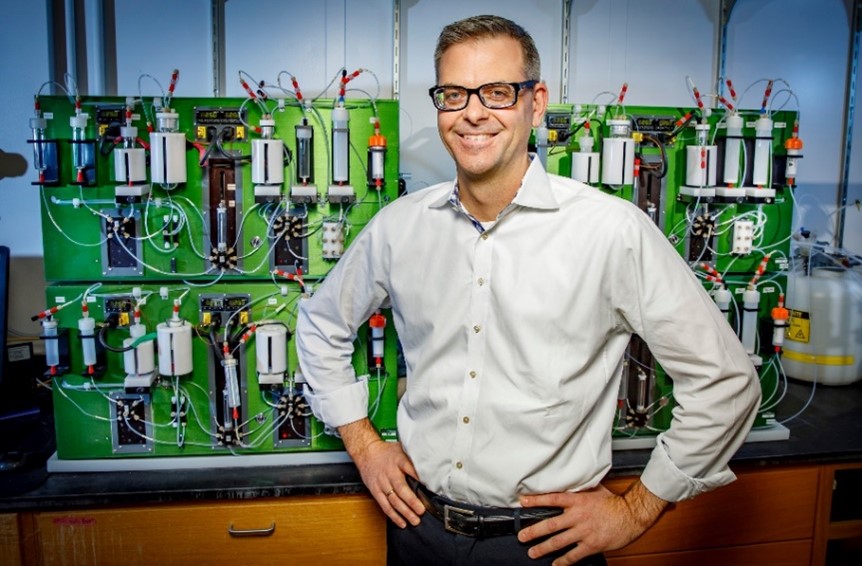

“At one point in the conversation, she stopped me and said, ‘It sounds like you know exactly what’s wrong with me, so why can’t you fix it?’” Dr. Burke, now the May and Ving Lee Professor for Chemical Innovation at University of Illinois Urbana-Champaign (UIUC), never forgot this question. In fact, it’s inspired his career-long search for new ways to develop therapies for diseases without effective treatment options.
Continue reading “Martin Burke: Replacing Lost Proteins to Treat Disease”