To dismantle the viruses that infect them, bacteria have evolved an immune system that identifies invading viral DNA and signals for its destruction. This gene-editing system is called CRISPR, and it’s being harnessed as a tool for modifying human genes associated with disease.
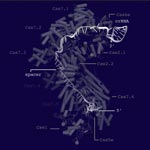
Taking another important step toward this potential application, researchers now know the structure of a key CRISPR component: a multi-subunit surveillance machine called Cascade that identifies the viral DNA. Shaped like a sea horse, Cascade is composed of 11 proteins and CRISPR-related RNA. A research team led by Blake Wiedenheft of Montana State University used X-ray crystallography and computational analysis to determine Cascade’s configuration. In a complementary study, Scott Bailey of Johns Hopkins University and his colleagues determined the structure of the complex bound to a viral DNA target.
Like blueprints, these structural models help scientists understand how Cascade assembles into an efficient surveillance machine and, more broadly, how the CRISPR system functions and how to adapt it as a tool for basic and clinical research.
Learn more:
Montana State University News Release ![]()
Johns Hopkins University News Release ![]()