What do you get when you mix a room full of scientists with a classroom full of students who have questions about cells? Cell Day 2016! During this free web chat, middle and high school students will have the opportunity to ask our scientists at NIGMS about cell biology, biochemistry, research careers and more. Join us on Thursday, November 3 anytime from 10 a.m. to 3 p.m. EDT. Registration (no longer available).
Posts by Shelly Pollard
Nanoparticles Developed to Stick to Damaged Blood Vessels, Deliver Drugs
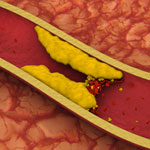
Heart disease is the leading cause of death for both men and women in the United States, according to the Centers for Disease Control and Prevention. One treatment challenge is developing non-invasive ways to direct medication to damaged or clogged arteries, which can block blood flow and increase the risk for heart attack and stroke. A team led by Naren Vyavahare at Clemson University has engineered extremely tiny particles—nanoparticles—that offer a promising step forward.
Healthy arteries have elastic fibers that make the arteries flexible. But, in most cardiovascular diseases, the fibers get damaged. The new nanoparticles, which can deliver drugs, attach only to damaged fibers and could enable site-specific drug delivery to minimize off-target side effects. The nanoparticles also allow drugs to be released over longer periods of time, potentially increasing the drugs’ effectiveness. The new biomaterial was tested in rodent models for studying vascular disease, so it is still in the early stages of development.
This work also was funded by NIH’s National Heart, Lung, and Blood Institute.
Dendrites Show Ability to Regenerate After Injury
When a bone breaks, it might slice axons—the part of nerve cells that sends information to other cells—and potentially cause loss of mobility or feeling. Prior research had shown that a damaged nerve cell could repair such an injury through the regrowth of axons. Scientists at Penn State University wondered if dendrites—the part of nerve cells that receive information from other nerve cells—could also regenerate. To find out, Melissa Rolls and her team cut off the dendrites from nerve cells in fruit flies. Instead of dying, as was expected, the cells regrew dendrites. The research also revealed that dendrite regeneration happens independently of axon regeneration, leading investigators to believe there are two separate regeneration pathways: one for axons and one for dendrites. Learning more about this new dendrite regrowth pathway might one day lead to new approaches for healing injured nerve cells, including those damaged after a stroke.
Learn more:
Penn State News Release ![]()
Rolls Lab
Transporter Protein Dance Moves
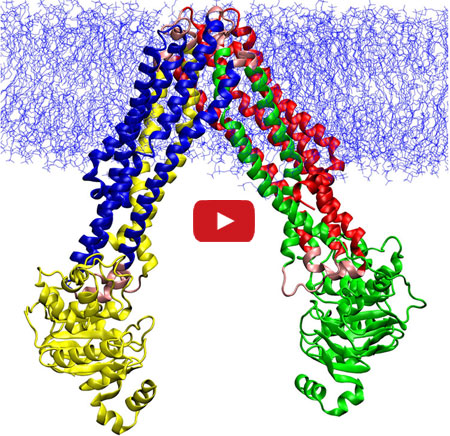
In this video, Emad Tajkhorshid of the University of Illinois at Urbana-Champaign explains the molecular dance of ABC transporters, a family of molecular machines that utilize ATP to move substances across the cell membrane. Tajkhorshid and his team recently used computational methods to map the movements between two known structural models of MsbA, a bacterial version of a transporter in human cells that helps to export anti-cancer drugs. They then described the individual steps of the molecular motions during the transport cycle. Understanding the process at such a detailed level could suggest new targets for treating a range of diseases, including some drug-resistant cancers that often make more transporter proteins to kick out medications meant to kill them.
Learn more:
University of Illinois at Urbana-Champaign News Release ![]()
Tajkhorshid Lab
Abnormal Mitochondria Might Cause Resistance to Radiation Therapy

Why some cancers are resistant to radiation therapy has baffled scientists, but research on abnormalities in mitochondria, often described as cells’ power plants, could offer new details. A research team led by Maxim Frolov ![]() of the University of Illinois at Chicago learned that the E2F gene, which plays a role in the natural process of cell death, contributes to the function of mitochondria. Fruit flies with a mutant version of the E2F gene had misshapen mitochondria that produced less energy than normal ones. Flies with severely damaged mitochondria were more resistant to radiation-induced cell death. Studies using human cells revealed similar effects. The work could help explain why people with cancer respond differently to radiation therapy and might aid the development of drugs that enhance mitochondrial function, thereby improving the effectiveness of radiation therapy.
of the University of Illinois at Chicago learned that the E2F gene, which plays a role in the natural process of cell death, contributes to the function of mitochondria. Fruit flies with a mutant version of the E2F gene had misshapen mitochondria that produced less energy than normal ones. Flies with severely damaged mitochondria were more resistant to radiation-induced cell death. Studies using human cells revealed similar effects. The work could help explain why people with cancer respond differently to radiation therapy and might aid the development of drugs that enhance mitochondrial function, thereby improving the effectiveness of radiation therapy.
This work also was funded by NIH’s National Cancer Institute.
Learn more:
University of Illinois at Chicago News Release ![]()
New Approach Subtypes Cancers by Shared Genetic Effects

Cancer tumors are like snowflakes—no two ever share the same genetic mutations. Their unique characteristics make them difficult to categorize and treat. A new approach proposed by Trey Ideker and his team at the University of California, San Diego, might offer a solution. Their approach, called network-based stratification (NBS), identifies cancer subtypes by how different mutations in different cancer patients affect the same biological networks, such as genetic pathways. As proof of principle, they applied the method to ovarian, uterine and lung cancer data to obtain biological and clinical information about mutation profiles. Such cancer subtyping shows promise in helping to develop more effective, personalized treatments.
Learn more:
University of California, San Diego News Release
Ideker Lab
How Some Bacteria Colonize the Gut
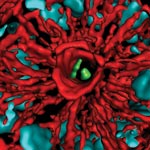
Have you ever felt that your gut was trying to tell you something? The guts of germ-free mice have told scientists a few new things about our resident microorganisms. By studying a genus of bacteria called Bacteriodes that live in the gastrointestinal tract, Sarkis Mazmanian of the California Institute of Technology discovered how Bacteriodes species stake their claim in a mouse’s gut. Mazmanian and his colleagues introduced different species of Bacteriodes into germ-free mice to learn how the microbes competed and found that most of them peacefully co-existed. However, when microbes of a species that was already present were introduced, they couldn’t take up residence. Further investigation revealed that Bacteriodes species, due to a set of specific genes, can live in tiny pockets or “crypts” of the colon, where they are sheltered from antibiotics and infectious microbes passing through. Understanding how these microorganisms colonize could help devise ways to correct for abnormal changes in bacterial communities that are associated with disorders like inflammatory bowel disease.
This work also was funded by NIH’s National Institute of Diabetes and Digestive and Kidney Diseases.
Learn more:
California Institute of Technology News Release
Mazmanian Lab
Mazmanian Video Interview ![]()
Gut Microbes Can Inactivate Cardiac Drugs
Doctors have known that a medication often prescribed to treat heart failure is inactivated by gut microbes, particularly a bacterial species called E. lenta. Now scientists have a better understanding of why. A Harvard University research team led by Peter Turnbaugh found that the heart drug digoxin turns on two E. lenta genes that help convert the drug into its inactive form, thereby making the medicine less effective. By measuring gene abundance, the scientists could reliably predict whether a microbial community could break down the drug. The researchers also identified a possible way to stop the process: add protein. Their studies using mice showed that a diet high in protein—and the amino acid, arginine, that helps E. lenta grow—increased digoxin absorption. These initial findings suggest that one day it may be possible to tailor digoxin therapy through diet modifications.
Learn more:
Harvard University News Release ![]()
Turnbaugh lab ![]()

