We asked the heads of our scientific divisions to tell us about some of the big questions in fundamental biomedical science that researchers are investigating with NIGMS support. This article is the third in an occasional series that explores these questions and explains how pursuing the answers could advance understanding of important biological processes.
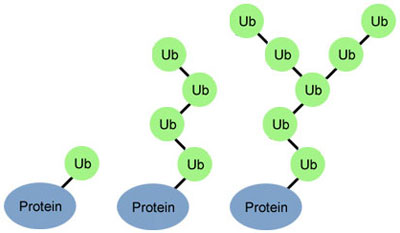
Researchers are on a quest to crack a code made by ubiquitin, a small protein that plays a big role in coordinating cellular function. By attaching to other proteins, ubiquitin determines what those proteins should do next.
Just as zip codes direct letters to specific towns, the ubiquitin code might direct one protein to help with DNA repair, another to assist in cell division, and a third to transport molecules into and out of cells.
The ubiquitin code is, well, ubiquitous. It drives basic cellular processes in organisms as diverse as yeast, plants and humans. Yet despite the code’s prevalence, scientists still have much to learn about how it works. A big challenge is figuring out how the shapes of what might be hundreds of different ubiquitin molecules drive distinct protein actions. There’s also evidence that a single ubiquitin formation can code for multiple different protein fates.
Robert Cohen of Colorado State University and his team are helping solve the ubiquitin puzzle by developing sensors that allow scientists to watch as ubiquitin guides cellular processes in live cells.
Their strategy works like this: First, they choose a ubiquitin complex with a known shape that attaches to a protein whose shape is also known. Then they design a sensor molecule that should attach to points on both the ubiquitin complex and the protein.
Next, in test tubes they measure this sensor’s fit and tweak its shape until it attaches tightly to the ubiquitin-protein pair. Finally, they introduce it into live cells.
The sensor typically contains a fluorescent protein that allows the scientists to view its movement in the cell. Wherever they see this sensor glow, they know that the ubiquitin-protein pair is present. By watching the pair as cells grow or undergo stress, the scientists can gain a better sense of which cellular process the ubiquitin formation helps guide.
The team is currently using one sensor to study how ubiquitin in human cells regulates the response to DNA damage.
In addition to determining the function of ubiquitin formations paired with proteins, these sensors may shed light on ubiquitin-related conditions.
“Many human conditions such as cancers as well as neurodegenerative and developmental disorders develop when cells misread ubiquitin signals. Scientists are working hard to design therapies to fix these errors,” Cohen says. “We hope that our imaging tools will one day allow others to evaluate how drugs affect the locations and concentrations of ubiquitin in real time in living cells.”
This research was funded in part by NIH grants R01GM097452 and R01GM115997.

