
Credit: Wikimedia Commons, Usman Bashir.
Dictyostelium discoideum
Natural habitat: Deciduous forest soil and moist leaf litter
Favorite food: Bacteria
Top speed: 8 micrometers per minute
Like the athletes in Rio, the world’s most highly advanced microbial runners recently gathered in Charlestown, Massachusetts, to find out which ones could use chemical cues to most quickly navigate a maze-like microfluidic racecourse. The winners’ prize: credit for helping scientists learn more about how immune system cells navigate through the human body on their way to fight disease.
The finalists were a group of soil-dwelling slime molds called Dictyostelium that were genetically engineered by a pair of Dutch biochemists to detect minuscule chemical changes in the environment. The racers used their enhanced sense of “smell” to avoid getting lost on their way to the finish line.
While researchers have been racing the genetically souped-up microbes at annual events for a few years—another competition is scheduled for October 26—scientists have been studying conventional Dictyostelium for decades to investigate other important basic life processes including early development, gene function, self/non-self recognition, cell-type regulation, chemical signaling and programmed cell death.
We “interviewed” a member of the winning team, who calls herself Dicty, to hear more about the many important contributions Dictyostelium make to science. “People get a little grossed out when I mention that I’m a slime mold,” says Dicty. “But I’m versatile and really easy to work with.”
Dictyostelium work their way through a microfluidic maze by following chemical attractants. Credit: Daniel Irimia, Massachusetts General Hospital.
A Star is Born
Dictyostelium first grabbed the public’s imagination in the 1950s when long-time slime mold researcher John Bonner shot grainy black and white videos  illustrating the creatures’ complicated, but striking life cycle. In the videos, solitary unicellular Dictyostelium jitterbug around nibbling stray bacteria. Suddenly, when their food supply dries up, more than 100,000 swarm together and coalesce into a migratory slug, kind of like a microbial Voltron. They then stack on top of one another and form a long stalk topped with a round capsule full of spores called a fruiting body. Biologists investigate this process as a way to learn more about how unicellular organisms came together to form multicellular life.
illustrating the creatures’ complicated, but striking life cycle. In the videos, solitary unicellular Dictyostelium jitterbug around nibbling stray bacteria. Suddenly, when their food supply dries up, more than 100,000 swarm together and coalesce into a migratory slug, kind of like a microbial Voltron. They then stack on top of one another and form a long stalk topped with a round capsule full of spores called a fruiting body. Biologists investigate this process as a way to learn more about how unicellular organisms came together to form multicellular life.
The mobility of Dictyostelium makes it an exceptional model organism for studying the movement of neutrophils, commonly known as white blood cells. Neutrophils are our first line of defense against pathogens. They rush to the site of new wounds, kill bacteria and use chemical signals to initiate the body’s immune response. Learning about neutrophil migration is challenging because these cells are notoriously difficult to study in the lab. “We crawl around pretty much just like neutrophils, but we’re way easier to observe in Petri dishes,” says Dicty, adding, “or microfluidic racecourses.”
Time lapse of Dictyostelium colony in process of aggregation. Credit: John Bonner.
Scientists can investigate strains of Dictyostelium with genetic defects that affect their ability to navigate. Such research has revealed biochemical communication networks within Dictyostelium that are important for their migration and have shown that some of these same networks also guide movement in neutrophils.
The Versatile Model Organism
Dictyostelium’s swarming behavior has also inspired scientists to use the tiny critters as a model for allorecognition, an organism’s ability to distinguish its own cells from those of another. Genetic experiments show that Dictyostelium use two sets of proteins to recognize cells from the same strain. “If my proteins don’t match up with those of my neighbors, I’m definitely not going to join with them,” says Dicty. Understanding how this relatively simple organism tells friend from foe can illuminate the mechanisms our own immune systems use to identify invading pathogens.
Another intriguing feature of Dictyostelium that is obvious from Bonner’s videos is the continuous shape shifting they undergo as they meander. Researchers have used Dictyostelium to screen thousands of molecules for those that are responsible for altering cell shape. These molecules may one day provide clues for the development of therapies for conditions such as cancer and chronic obstructive pulmonary disease, which are characterized by changes in cell size and shape.
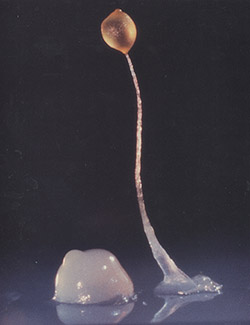
A group of 100,000 individual Dictyostelium come together to form a long stalk topped with a round capsule full of spores called a fruiting body. Credit: Rex Chisholm, Northwestern University.
A few years ago researchers added a surprisingly complex task to the list of things Dictyostelium can do: farming. About a third of Dictyostelium observed in the wild engage in the husbandry of bacteria. “When my friends and I notice we’re about to run out of food, we stop chowing down on bacteria and stow some away in our fruiting body for later,” says Dicty. Then, during spore dispersal, Dictyostelium carry their store of bacteria with them and use it to seed a new food crop. This can be a major advantage if the spores land in an area lacking edible bacteria.
Bacteria stored in fruiting bodies lend another important advantage to Dictyostelium trying to take root in a new location: protection from environmental toxins. Researchers speculate that the bacteria excrete chemicals to degrade deadly toxins or act as a sponge to soak them up, shielding Dictyostelium from harm. Scientists hope that exploring the relationship between Dictyostelium and bacteria may help reveal novel ways that the billions of microbes inhabiting humans protect their hosts from toxins they encounter.
“It’s always a bit amusing when people get excited about some ‘new’ thing we can do,” says Dicty. “I’m just tickled that scientists continue to find ways for me and my fellow Dictyostelium to contribute to the advancement of human health and knowledge of the world.”
Portions of this research were funded by NIH under grants R01GM092804, R01GM066817 and R01GM084992.


![]() illustrating the creatures’ complicated, but striking life cycle. In the videos, solitary unicellular Dictyostelium jitterbug around nibbling stray bacteria. Suddenly, when their food supply dries up, more than 100,000 swarm together and coalesce into a migratory slug, kind of like a microbial Voltron. They then stack on top of one another and form a long stalk topped with a round capsule full of spores called a fruiting body. Biologists investigate this process as a way to learn more about how unicellular organisms came together to form multicellular life.
illustrating the creatures’ complicated, but striking life cycle. In the videos, solitary unicellular Dictyostelium jitterbug around nibbling stray bacteria. Suddenly, when their food supply dries up, more than 100,000 swarm together and coalesce into a migratory slug, kind of like a microbial Voltron. They then stack on top of one another and form a long stalk topped with a round capsule full of spores called a fruiting body. Biologists investigate this process as a way to learn more about how unicellular organisms came together to form multicellular life.


Nice information sir