
“Being a scientist is thrilling, and it’s also tremendously fun,” says Eda Koculi, Ph.D., assistant professor of chemistry and biochemistry at the University of Texas at El Paso (UTEP). “In my opinion, science is the only profession that allows a person to simultaneously express their creativity, quench their intellectual curiosity, and serve society.” We spoke with Dr. Koculi about how she became a researcher, what she’s uncovering about how ribosomes are built and modified, and how she encourages students to pursue scientific careers.
Get to Know Dr. Koculi
- Coffee or tea? Coffee
- Favorite music genre? Classical
- Salty or sweet? Salty
- Early bird or night owl? Night owl
- Washing glassware in the lab or dishes in your kitchen? Glassware
- What was your childhood dream job? A scientist or a teacher—and I have both my dream jobs.
- Favorite hobby? Hiking
- Favorite piece of lab safety equipment? Geiger counter
- Favorite molecule? RNA
- A scientist (past or present) you’d like to meet? Marie Curie
Continue reading “Exploring Ribosome Assembly and RNA Modification: Q&A With Eda Koculi”
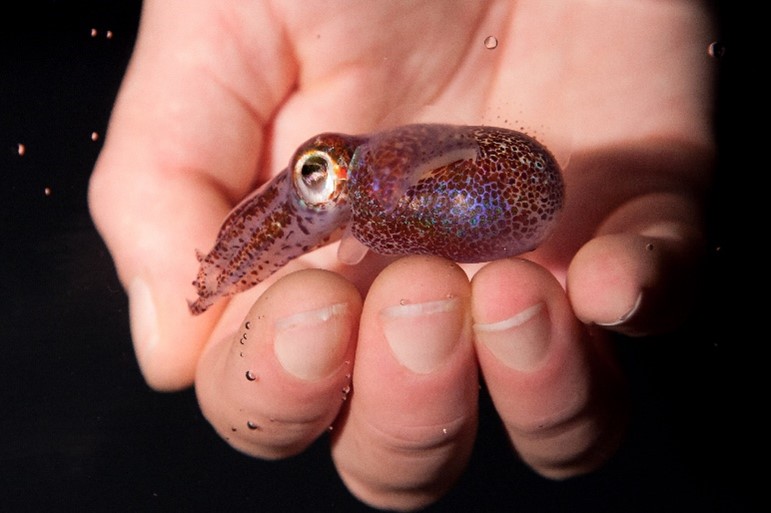
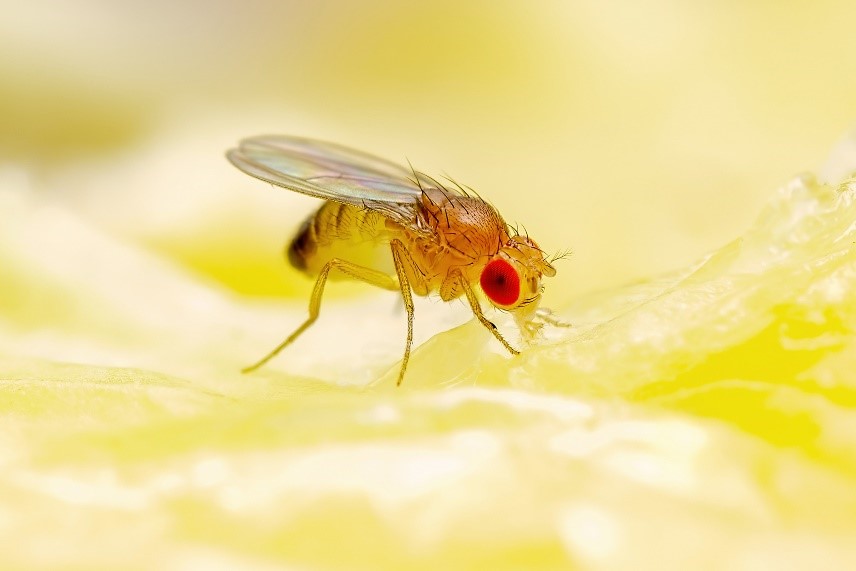
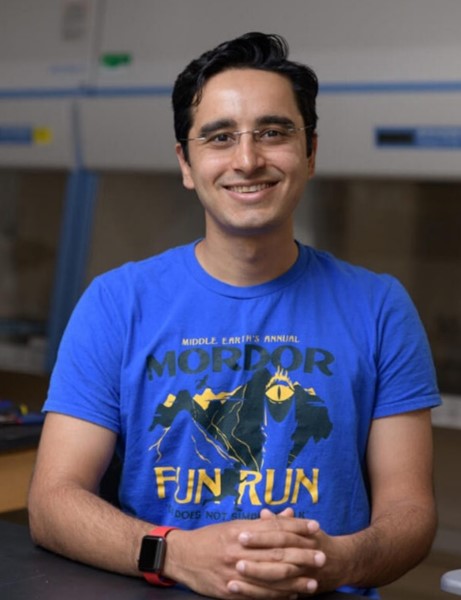
















 Credit: Adapted from Compound Interest.
Credit: Adapted from Compound Interest.