
In less than two decades, RNA interference (RNAi)—a natural process cells use to inactivate, or silence, specific genes—has progressed from a fundamental finding to a powerful research tool and a potential therapeutic approach. To check in on this fast-moving field, I spoke to geneticists Craig Mello of the University of Massachusetts Medical School and Michael Bender of NIGMS. Mello shared the 2006 Nobel Prize in physiology or medicine with Andrew Fire ![]() of Stanford University School of Medicine for the discovery of RNAi. Bender manages NIGMS grants in areas that include RNAi research.
of Stanford University School of Medicine for the discovery of RNAi. Bender manages NIGMS grants in areas that include RNAi research.
How have researchers built on the initial discovery of RNAi?
A scientific floodgate opened after the 1998 discovery that it was possible to switch off specific genes by feeding microscopic worms called C. elegans double-stranded RNA that had the same sequence of genetic building blocks as a target gene. (Double-stranded RNA is a type of RNA molecule often found in, or produced by, viruses.) Scientists investigating gene function quickly began to test RNAi as a gene-silencing technique in other organisms and found that they could use it to manipulate gene activity in many different model systems. Additional studies led the way toward getting the technique to work in cells from mammals, which scientists first demonstrated in 2001. Soon, researchers were exploring the potential of RNAi to treat human disease.
What do we know now about how RNAi works?
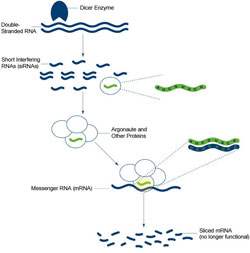
In the first step of RNAi, an enzyme named Dicer chews up the double-stranded RNA into smaller RNA pieces known as short interfering RNAs (siRNAs). The cell’s RNAi machinery uses these snippets of RNA as guides for finding matching genetic sequences in messenger RNA (mRNA) molecules, which transmit information coded in genes to the cell’s protein factories. The machinery then slices these mRNAs, effectively destroying their messages and silencing the corresponding gene. Proteins called Argonautes are the key facilitators of RNAi’s search-and-destroy process in worms as well as in humans.
Why does RNAi exist?
Cells in organisms including C. elegans and plants use RNAi to find and eliminate invading viruses. Human cells contain all the necessary RNAi components to do the same, but the jury is still out on whether they use an RNAi-like response to control viruses. Our cells do use RNAi to control the spread of genetic elements called transposons that can jump from place to place in the genome and create genetic changes that can lead to cancer or other diseases.
How is RNAi advancing biomedical research?
Scientists are using RNAi as a powerful research tool to help figure out how cells work. In the lab, investigators are harnessing RNAi to turn off any gene in a cell and answer questions about gene function that were once well beyond their reach. The technique is speeding the discovery of key players in normal and disease processes, allowing scientists to identify many new targets for drug development.
What attributes of RNAi make it attractive as an approach to treating human disease?
In principle, the RNAi approach should work for virtually any gene. The disorders most likely to respond to RNAi-based therapies are ones that can be treated by knocking down the activity of one or several genes involved in disease. By delivering custom-made siRNA molecules to the desired target in the body, scientists can employ the built-in components of the cell’s own RNAi machinery to seek out and silence specific disease genes.
Furthermore, many current medicines work by targeting enzymes—proteins that speed up chemical reactions in our cells—but most proteins lack enzymatic activities and therefore are difficult or impossible to target. An RNAi-based approach gets around this problem by preventing a gene from producing a disease-causing or malfunctioning protein in the first place.
How is research on RNAi realizing its potential for improving human health?
Today, researchers are testing RNAi-based therapies for diseases ranging from cancer and macular degeneration to a rare, inherited liver disease called TTR-mediated amyloidosis. Viral infections are also potential targets for RNAi-based therapies. Although no RNAi-based therapies have been approved for use as of now, roughly two dozen are in clinical trials in humans.
What challenges still exist for the use of RNAi in the clinic?
The major obstacle has been delivering RNAi-based therapies to the right cell, tissue or organ in the body. Another challenge is preventing unwanted side effects that can occur when therapeutic siRNAs or the molecules used to deliver them trigger an immune response. Also, RNAi-based therapies may need to be given either intravenously or by injection and over a long period; they are not a permanent fix.
How are NIGMS-funded researchers and others working to overcome these challenges?
Researchers have developed different ways of delivering siRNAs to the desired cells, tissues or organs and are continuing to refine these methods. The strategies involve packaging siRNA molecules in tiny carrier particles or chemically modifying them. These approaches can make the siRNAs more stable, let them enter cells more readily and reduce side effects. Researchers can also attach siRNAs to other molecules, such as antibodies, that selectively target the therapeutic siRNA to diseased cells or tissues.
So far, these methods have been most effective for delivering siRNAs to a limited number of tissues, including the liver. As a result, some of the more promising clinical trials of RNAi are targeting liver-linked diseases like amyloidosis. Injecting chemically modified siRNAs directly into the eye or skin has been effective, too, and early-phase clinical trials of RNAi-based therapies are in progress for several eye and retinal disorders as well as for reducing post-surgery skin scarring.
Scientists are also studying delivery strategies that involve splicing the genetic instructions for making the siRNA into a genetically hobbled carrier virus. Theoretically, once the modified virus infects a cell, the cell will produce the siRNA indefinitely, eliminating the need for repeated treatments.
Other researchers continue to explore the basic biology of RNAi and dissect its actions step-by-step. Insights from this work could further fuel progress toward developing more and better RNAi-based therapies.


Excellent article. Please keep us informed and up-to-date. I hope practicing doctors treating patients are made aware of the available treatmentss as they become available.
Very thorough and easy to understand article.
RNA Interference research has a lot of potential to be utilized within the context of ‘Precision Medicine’. With the exponential evolution of Genomic data showing us that many mutations showing variances of unknown significance may be an area this concept might have some traction in the future..