RNA is essential for life as we know it. Among other roles, this molecule helps translate the instructions of DNA into proteins, which perform a vast range of tasks to keep us alive and healthy. In past Biomedical Beat posts, we’ve discussed the basics of RNA and how researchers are using it to develop medicines, vaccines, and tests for certain diseases. This year, in honor of RNA Day on August 1, we’ve created a quiz all about this remarkable molecule. Test your knowledge with the questions below!
Continue reading “Quiz: Can You Solve These RNA Riddles?”Tag: RNA
Exploring Ribosome Assembly and RNA Modification: Q&A With Eda Koculi

“Being a scientist is thrilling, and it’s also tremendously fun,” says Eda Koculi, Ph.D., assistant professor of chemistry and biochemistry at the University of Texas at El Paso (UTEP). “In my opinion, science is the only profession that allows a person to simultaneously express their creativity, quench their intellectual curiosity, and serve society.” We spoke with Dr. Koculi about how she became a researcher, what she’s uncovering about how ribosomes are built and modified, and how she encourages students to pursue scientific careers.
Get to Know Dr. Koculi
- Coffee or tea? Coffee
- Favorite music genre? Classical
- Salty or sweet? Salty
- Early bird or night owl? Night owl
- Washing glassware in the lab or dishes in your kitchen? Glassware
- What was your childhood dream job? A scientist or a teacher—and I have both my dream jobs.
- Favorite hobby? Hiking
- Favorite piece of lab safety equipment? Geiger counter
- Favorite molecule? RNA
- A scientist (past or present) you’d like to meet? Marie Curie
Continue reading “Exploring Ribosome Assembly and RNA Modification: Q&A With Eda Koculi”
Understanding RNA-Modifying Enzymes: Q&A With Jeffrey Mugridge

“One of the best aspects of research is the excitement of discovery, being the first person in the world to know a small detail about the system you’re studying,” says Jeffrey Mugridge, Ph.D., an assistant professor of chemistry and biochemistry at the University of Delaware in Newark. We talked with Dr. Mugridge about how a pet store job sparked his early interest in science, why he decided to change his career trajectory after graduate school, and what he believes is key to being a successful researcher.
Q: How did you first become interested in science?
A: My strong interest in science didn’t develop until I was in high
school—I wasn’t one of those kids who had a chemistry set or a deep love for dinosaurs or anything like that. But in high school, I worked in a pet store, where I learned a lot about aquarium science, including the ins and outs of managing water chemistry to keep fish alive. I also had a fantastic chemistry teacher who really helped me foster a love for the field.
Celebrating 10 Years of Biomedical Beat
This August marks 10 years of the blog! Throughout the past decade, we’ve brought you blog posts that explore basic science topics, quiz your knowledge, showcase cool images, and more! Some of our most-read favorites include:
Continue reading “Celebrating 10 Years of Biomedical Beat“Investigating the Secrets of Cancer-Causing Viruses

While she was in graduate school, Mandy Muller, Ph.D., became intrigued with viruses that are oncogenic, meaning they can cause cancer. At the time, she was researching human papillomaviruses (HPVs), which can lead to cervical and throat cancer, among other types. Now, as an assistant professor of microbiology at the University of Massachusetts (UMass) Amherst, Dr. Muller studies Kaposi sarcoma-associated herpesvirus (KSHV), which causes the rare AIDS-associated cancer Kaposi sarcoma.
A Continental Change
Dr. Muller has come a long way, both geographically and professionally, since her childhood in France. She was the first person in her family to graduate from high school, where she excelled in science, and went on to attend École Normale Supérieure (ENS) de Lyon, a research-oriented undergraduate institution in Lyon, France. “We spent weeks at a time in laboratory-based classes, working in real labs. That’s when I realized people could do research full-time, which caught my attention,” says Dr. Muller. She double-majored in biology and geology, and soon chose to focus her career on immunology and virology.
Continue reading “Investigating the Secrets of Cancer-Causing Viruses”In Other Words: Not All Bases Are in the Ballpark
You might first think about sports when you hear the word base, but not all bases are on the baseball diamond. In chemistry, a base is a molecule that reacts with an acid, often by accepting a proton from the acid or from water. Baking soda and dish soap are common bases.

Science Snippet: RNA’s Remarkable Roles
RNA, though less well known than its cousin DNA, is equally integral to our bodies. RNA molecules are long, usually single-stranded chains of nucleotides. (DNA molecules are also made up of nucleotides but are typically double-stranded.) There are three major types of RNA, which are all involved in protein synthesis:
- Messenger RNA (mRNA) is complementary to one of the DNA strands of a gene and carries genetic information for protein synthesis to the ribosome—the molecular complex in which proteins are made.
- Transfer RNA (tRNA) works with mRNA to make sure the right amino acids are inserted into the forming protein.
- Ribosomal RNA (rRNA), together with proteins, makes up ribosomes and functions to recognize the mRNA and tRNA that are presented to the ribosomal complex.
In Other Words: Translation Isn’t Only for Languages
In everyday use, most people understand translation to mean converting words from one language to another. But when biologists talk about translation, they mean the process of making proteins based on the genetic information encoded in messenger RNA (mRNA). Proteins are essential for virtually every process in our bodies, from transporting oxygen to defending against infection, so translation is vital for keeping us alive and healthy.

Scientist Interview: Exploring the Promise of RNA Switches with Christina Dawn Smolke
Whether animals are looking for food or mates, or avoiding pathogens and predators, they rely on biosensors—molecules that allow them to sense and respond to their environments. Christina Dawn Smolke, Ph.D., a professor of bioengineering at Stanford University in California, focuses her research on creating new kinds of biosensors to receive, process, and transmit molecular information. Her lab has built RNA molecules, or switches, that can alter gene expression based on biochemical changes they detect.
Continue reading “Scientist Interview: Exploring the Promise of RNA Switches with Christina Dawn Smolke”RNA Polymerase: A Target for New Antibiotic Drugs?
DNA, with its double-helix shape, is the stuff of genes. But genes themselves are only “recipes” for protein molecules, which are molecules that do the real heavy lifting (or do much of the work) inside cells.
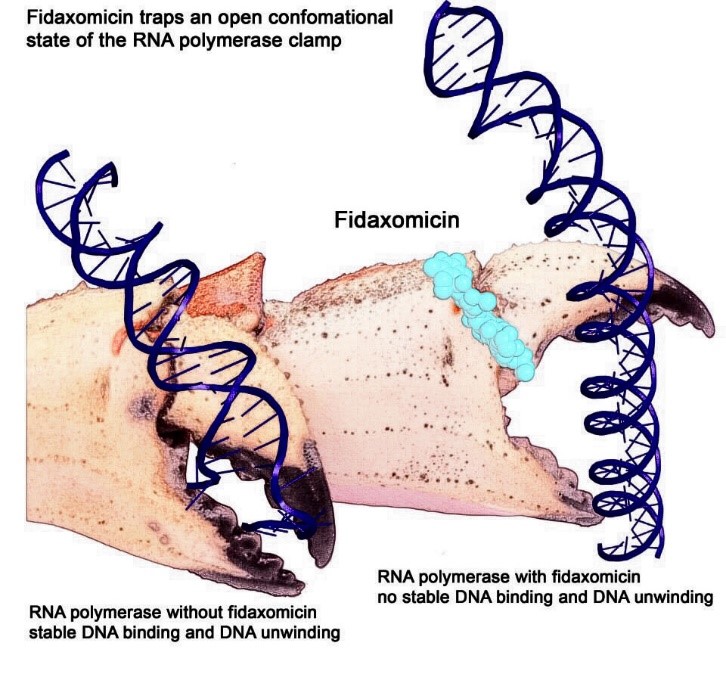 Artist interpretation of RNAP grasping and unwinding a DNA double helix. Credit: Wei Lin and Richard H. Ebright.
Artist interpretation of RNAP grasping and unwinding a DNA double helix. Credit: Wei Lin and Richard H. Ebright.Here’s how it works. A molecular machine called RNA polymerase (RNAP) travels along DNA to find a place where a gene begins. RNAP uses a crab-claw-like structure to grasp and unwind the DNA double helix at that spot. RNAP then copies (“transcribes”) the gene into messenger RNA (mRNA), a molecule similar to DNA.
The mRNA molecule travels to one of the cell’s many protein-making factories (ribosomes), which use the mRNA message as instructions for making a specific protein.
Continue reading “RNA Polymerase: A Target for New Antibiotic Drugs?”