To celebrate the 2024 National Postdoc Appreciation Week, we’re revisiting some scientists we’ve interviewed on the blog and how their postdoctoral experiences and NIGMS-funded training shaped their careers.



Biomedical Beat Blog – National Institute of General Medical Sciences
Follow the process of discovery
To celebrate the 2024 National Postdoc Appreciation Week, we’re revisiting some scientists we’ve interviewed on the blog and how their postdoctoral experiences and NIGMS-funded training shaped their careers.

Immunology is the study of the immune system, including all the cells, tissues, and organs that work together to protect you from germs. A person who studies immunology is called an immunologist, and there are three types:
The NIGMS Science Education Partnership Award (SEPA) program provides opportunities for pre-K-12 students from underserved communities to access STEM educational resources. SEPA grants support innovative, research-based, science education programs, furthering NIGMS’ mission to ensure a strong and diverse biomedical research workforce. SEPA projects generate resources that are mapped to state and national teaching standards for STEM and are rigorously evaluated for effectiveness; most are also available at no cost. These resources include mobile laboratories, interactive health exhibits in museums and science centers, educational resources for students, and professional development for teachers. Projects engage students and encourage them to envision themselves having careers in biomedical research.
To celebrate National STEM Day, we’re taking a look back at some of the SEPA projects we’ve recently featured on the blog, as well as our STEM teaching resources website, which includes several SEPA-funded materials. Check out the snapshots of each of the projects with links to the full articles and the teaching website below.
Continue reading “Spotlighting SEPA for National STEM Day”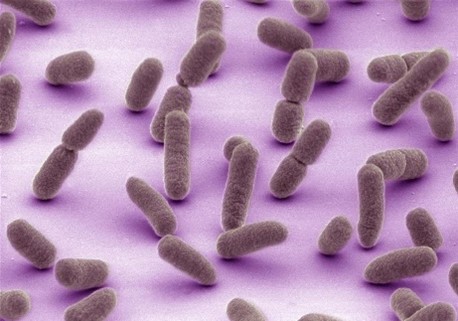
At least 1.7 million adults in the United States develop a life-threatening condition called sepsis each year. Sepsis is an overwhelming or impaired whole-body immune response that’s most often caused by bacterial infections. However, it can also be caused by viral infections, such as COVID-19 or influenza; fungal infections; or other injuries, including physical trauma. Anyone can get sepsis, but there’s a higher risk for some people, such as those who are ages 65 and older, who have certain medical conditions, or who have recently experienced severe illness or hospitalization.
The early symptoms of sepsis can include fever, chills, rapid breathing or heart rate, disorientation, and clammy or sweaty skin. Because other conditions also have these symptoms, sepsis can be difficult to diagnose. NIGMS-supported researchers are working to increase our understanding of sepsis so that doctors can identify it more quickly and treat it more effectively.
Continue reading “Quiz: Sepsis Science”This August marks 10 years of the blog! Throughout the past decade, we’ve brought you blog posts that explore basic science topics, quiz your knowledge, showcase cool images, and more! Some of our most-read favorites include:
Continue reading “Celebrating 10 Years of Biomedical Beat“Mentoring is a vital part of training the next generation of scientists. Through a variety of programs ranging from the undergraduate to faculty levels, NIGMS fosters the training and the development of a strong and diverse biomedical research workforce.
To celebrate National Mentoring Month, we’re highlighting a few of the many NIGMS-funded researchers who emphasize being great mentors. Check out the snapshots of our interviews with these mentors to see what they think about mentoring and to access and read their full stories.

Scientist Studies Burn Therapies After Being Severely Burned as a Child
Julia Bohannon, Ph.D., inspired by her own experience of being severely burned as a child, researches therapies that could prevent patients with burns from developing infections. Dr. Bohannon also mentors students, particularly those who hope to be both parents and scientists. “I’ve had a lot of women ask me for advice on how to be a mom and pursue a career in academia, and it’s been a really cool experience to be able to share that with students and trainees,” she says.
 Credit: ACS Website.
Credit: ACS Website.
It’s almost National Chemistry Week (NCW)! Each year, the American Chemical Society (ACS) unites scientists, undergraduate students, high school chemistry clubs, and other groups through this community-based program to reach the public—especially elementary and middle school
students—with positive chemistry messages.
Sometimes we can be our own worst enemies without even realizing it. One devastating example is sepsis: our body’s overwhelming or impaired immune response to an insult—usually an infection or an injury to the body. According to the Centers for Disease Control and Prevention (CDC), sepsis affects at least 1.7 million people in the United States each year, and it can lead to tissue damage, organ failure, and death. (See our sepsis fact sheet for more information.)

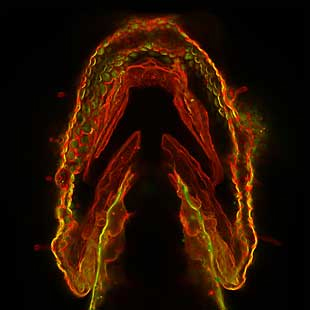
Large sugar molecules called glycans coat every cell in our bodies. They can also be found inside and between cells, and they are important for many biological processes, including how our cells interact with one another and with pathogens. For example, glycans on red blood cells determine blood type, and those on the cells of organs determine whether a person can receive a transplant from a particular donor. Scientists have only begun to explore sugars’ complexities and potential uses. Here, we look at the contributions three NIGMS-supported researchers are making to glycoscience.
Glycans called human milk oligosaccharides (HMOs) make up a significant portion of human milk. Study findings have shown that some HMOs can be prebiotics—substances that encourage beneficial bacteria to grow. Research has also revealed that some disease-causing microbes bind to certain HMOs, potentially allowing the germs to pass through the body without causing illness.
Continue reading “Could a Spoonful of Sugar Be a Medicine?”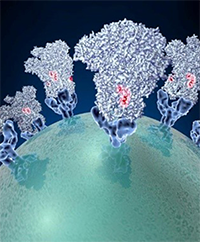 Spike proteins on the surface of a coronavirus. Credit: David Veesler, University of Washington.
Spike proteins on the surface of a coronavirus. Credit: David Veesler, University of Washington.
Since the start of the COVID-19 pandemic, researchers from many areas of biomedical science have worked together to learn how this new disease affects the human body, how to prevent its spread, and how to treat it. Severe cases of COVID-19 and cases of sepsis share many symptoms. Sepsis is the body’s overactive and extreme response to an infection. It’s unpredictable and can progress rapidly. Without prompt treatment, it can lead to tissue damage, organ failure, and death.
Sepsis has similarities with some cases of COVID-19, most likely because the two conditions trigger the same reactions at the cellular level. Researchers have studied these reactions in sepsis for many years.
“When we look back on 2020 and the speed with which progress was made against COVID-19, two features will stand out,” says John Younger, M.D., a member of the NIGMS Advisory Council who recently co-chaired a working group on advancing sepsis research. “The first is how quickly the biotechnology community came together to develop vaccine candidates. The second, and arguably the most immediately impactful, is how caregivers and clinical researchers were able to rapidly refine the care of COVID-19 patients based on decades of experience with sepsis.”
This post highlights a few of the many sepsis researchers supported by NIGMS who are applying their expertise to COVID-19.
Continue reading “Fight Against COVID-19 Aided by Sepsis Researchers”