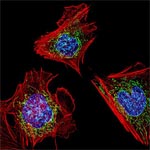
It might sound like a science fiction author made up genetic engineering, but it’s a real tool researchers use in the laboratory! A gene is a segment of DNA that codes for a protein. The information within a gene directs the building of a protein, block by block, through the process of gene expression. For a variety of reasons, including learning about certain cellular processes, scientists use genetic engineering in the lab to manipulate a cell’s genes and the proteins they encode.
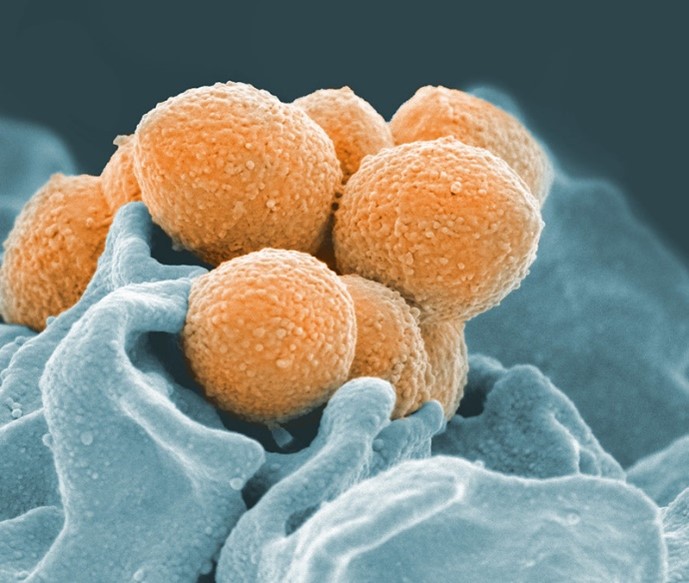
One of the most commonly used genetic engineering techniques is called clustered regularly interspaced short palindromic repeats (CRISPR), named for the odd, repeating sequences that researchers found in bacterial DNA in 1987.
Eventually, researchers discovered that these sequences are part of a bacterial immune system. (Just like humans, bacteria are susceptible to viral infections!) Some bacteria are able to insert short sequences of DNA from viruses that previously infected them into their own genome, allowing them to “remember” and more quickly recognize that virus in the future. If the invader tries to attack again, the bacterium recognizes and kills it by chopping up the part of its DNA that matches the “memory” using a special type of protein, an enzyme called CRISPR-Associated (Cas) protein. Our own immune systems also have the ability to remember pathogens through our adaptive immune response.
Continue reading “What Is CRISPR?”




 Jennifer Doudna, Ph.D. Credit: University of California, Berkeley.
Jennifer Doudna, Ph.D. Credit: University of California, Berkeley.