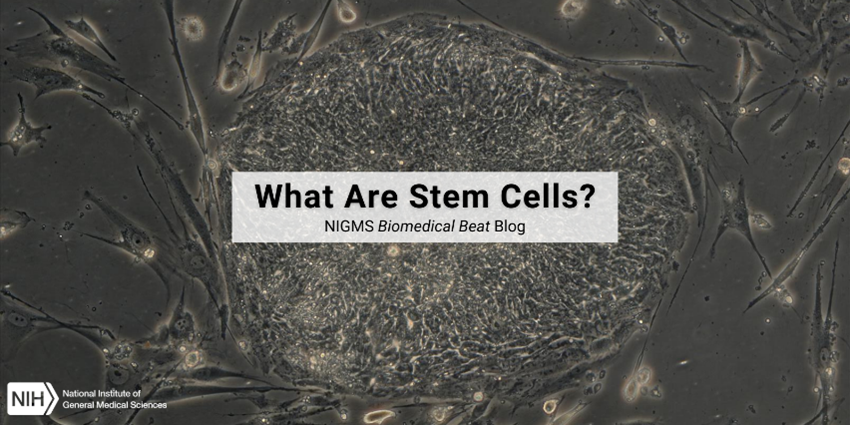
Stem cells are a special type of cell that is uniquely able to replicate many times and to develop into many different cell types. Stem cells live in tissues and organs throughout the body—including the liver, muscles, and skin—at all stages of life.
Continue reading “What Are Stem Cells?”Tag: Cells
Cells by the Numbers
Cells are the basic unit of life—and the focus of much scientific study. They’re categorized based on whether or not they have a distinct nucleus. Prokaryotic cells, like some bacteria such as blue-green algae, don’t have distinct nuclei. Instead, their nuclear material is spread throughout the cytoplasm. Eukaryotic cells—or cells with nuclei—make up humans, animals, plants, and fungi. Here are just a few of cells’ fascinating facets.
30 Trillion
That’s about how many human cells adults have in their bodies. Males are on the higher side with about 36 trillion cells, while females average about 28 trillion cells.
Continue reading “Cells by the Numbers”Membranes, Malaria, and the Mosaic of Science: Q&A With John Jimah

“I think it’s really an exciting time for science. Some people might think that everything out there to be discovered has already been discovered, but that’s far from the truth. There is still much, much more to discover,” says John Jimah, Ph.D., an assistant professor of molecular biology at Princeton University in Princeton, New Jersey. We talked with him about how he moved internationally to pursue his career, how his current research on cell membranes could help treat malaria, and how science holds space for everyone.
Get to Know Dr. Jimah
- Books or movies? Movies
- Coffee or tea? Mocha
- Beach or mountains? Beach
- Cats or dogs? Dogs
- Music, podcasts, or quiet? Podcasts
- Early bird or night owl? Early bird
- Childhood dream job? Judge
- Favorite hobby? Bicycling
- Favorite piece of lab safety equipment? Gloves
- A scientist (past or present) you’d like to meet? Leonardo da Vinci
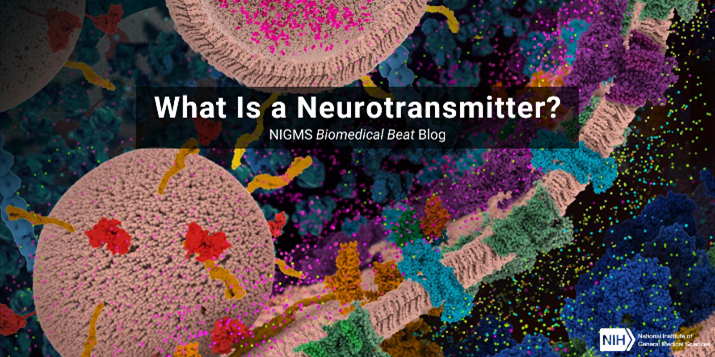
What Is a Neurotransmitter?
Have you wondered what controls the most basic functions of our bodies, like breathing, moving, and sleeping? Chemicals called neurotransmitters play a central role. Neurotransmitters pass messages from one nerve cell to another, and sometimes to muscles or glands. These messages may:
- Prompt the next nerve cell to pass on the message, prevent the message from going any further, or adjust how the message is passed on
- Cause a muscle to contract, like our intestines do when they digest food
- Tell a gland to secrete hormones, which are molecules that further pass on messages to tissues or organs
Quiz: Can You Solve These RNA Riddles?
RNA is essential for life as we know it. Among other roles, this molecule helps translate the instructions of DNA into proteins, which perform a vast range of tasks to keep us alive and healthy. In past Biomedical Beat posts, we’ve discussed the basics of RNA and how researchers are using it to develop medicines, vaccines, and tests for certain diseases. This year, in honor of RNA Day on August 1, we’ve created a quiz all about this remarkable molecule. Test your knowledge with the questions below!
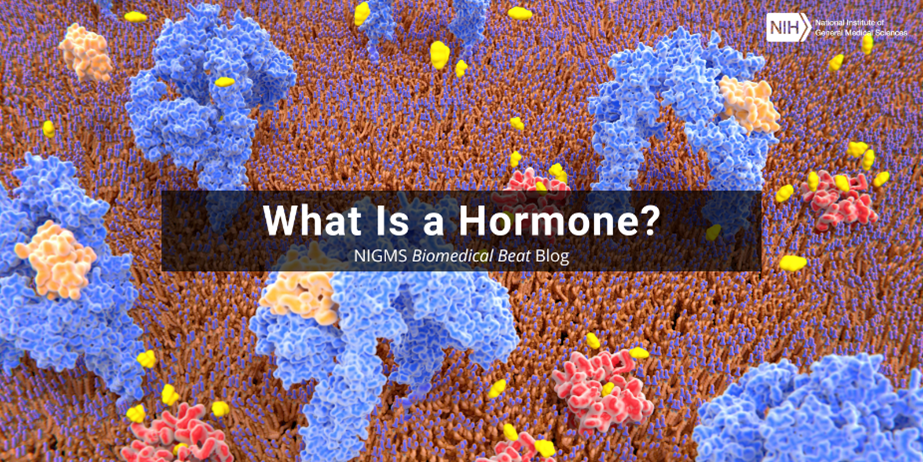
Continue reading “Quiz: Can You Solve These RNA Riddles?”What Is a Hormone?
Hormones are chemical messengers in the body that glands form and release, or secrete, into the bloodstream, where they travel to various organs and tissues to change biological functions. Hormone levels fluctuate during a lifespan and even on a daily basis.
Growth spurts in toddlers or sudden changes in adolescents are directly related to large hormonal shifts during development and puberty. Smaller changes occur throughout each day to help maintain normal bodily functions, such as our sleep-wake cycle known as our circadian rhythm.
Continue reading “What Is a Hormone?”The Third Product of Cell Division: Q&A With Ahna Skop

“Throughout my career, I’ve enjoyed studying topics that no one else seems to care about. I always tell people that I like searching through the scientific garbage bin for inspiration,” says Ahna Skop, Ph.D., a professor of genetics at the University of Wisconsin-Madison. We talked with her about the backyard experiment that helped her gain confidence in her scientific abilities, her career-long pursuit to better understand a detail about cell division that others had written off as unimportant, and her desire to build an accessible scientific community.
Q: How did you first become interested in science?
A: Middle school and high school science fairs had a big impact on me. I would develop my ideas, and with the help of my dad, build the experimental setup I needed to answer the scientific question. One of my experiments studied whether ants preferred to eat salt or sugar, so I poured small piles of both all over the backyard and took daily measurements of the height of the piles to figure out which type was shrinking faster. (Spoiler alert for those of you who might try this at home: They liked both but preferred the sugar to the salt.)
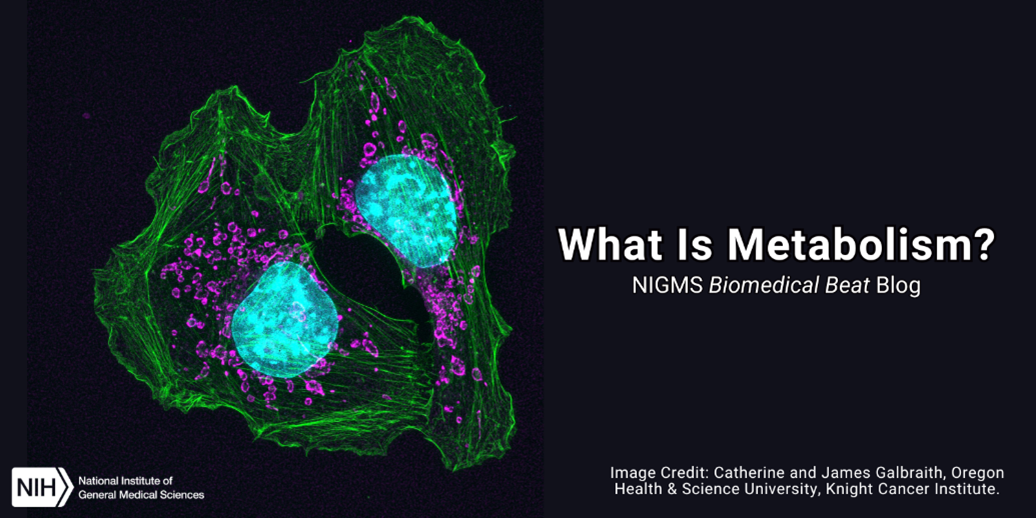
Continue reading “The Third Product of Cell Division: Q&A With Ahna Skop”What Is Metabolism?
You’ve likely heard someone attribute their body size to a fast or slow metabolism. But did you know there’s much more to metabolism than calories burned? Metabolism includes all the chemical changes that occur as our bodies use enzymes to break down food, medicines, and biological substances as well as produce energy and materials needed for growth.
The Two Sides of Metabolism
Our bodies have many metabolic pathways, but they all fall into two main categories: catabolic and anabolic. Catabolic pathways break down complex molecules into simpler ones, usually releasing energy in the process. For example, catabolic pathways turn large carbohydrate molecules from our food into simple sugars, such as glucose. Some of the most well-known catabolic pathways then convert the simple sugar glucose into adenosine triphosphate (ATP), a molecule that cells commonly use as an energy source.
Continue reading “What Is Metabolism?”Investigating the Primary Cilium: Q&A With Xuecai Ge

The brain is a large and complex organ, but some very small structures guide its development. Xuecai Ge, Ph.D., an associate professor of molecular and cell biology at the University of California, Merced (UC Merced), has devoted her career to understanding one of these structures called the primary cilium. In an interview, Dr. Ge shared how her childhood experience inspired her to study science and what makes the primary cilium fascinating.
Q: How did you first become interested in science?
A: When I was a little kid, my mom was a primary care doctor, and I saw her treat patients in our community. I noticed that no matter who got a particular illness, she could use the same medicine to treat them. My little mind was amazed that the same medicine could work for so many different people! I think this early experience planted the original seed of my interest in life science.
Continue reading “Investigating the Primary Cilium: Q&A With Xuecai Ge”Science Snippet: Examining Enzymes
Every day, our cells must produce all the various molecules they need to stay alive. But the chemical reactions to create these molecules can’t occur without help—which is where enzymes come in. Enzymes are biological catalysts, meaning they speed up the rate of specific chemical reactions by reducing the amount of energy needed for the reaction to occur. Most enzymes are proteins, but some RNA molecules can also act as enzymes.
Thousands of different enzymes catalyze the vast range of reactions that take place within cells, but each enzyme typically supports one of the following types of tasks:
Continue reading “Science Snippet: Examining Enzymes”