 Protein Data Bank’s 50 years logo. Credit: PDB website.
Protein Data Bank’s 50 years logo. Credit: PDB website.
The Protein Data Bank (PDB), established in 1971, is the single global repository for 3D structural data of proteins, DNA, RNA, and even complexes these biological molecules form with drugs or other small molecules. More than 1 million people—including researchers, medical professionals, educators, and students—use the PDB each year. NIGMS and other parts of NIH have helped fund this free digital resource since 1978.
Simulate Scientific Discovery & Learn the Steps to Unlocking Molecular Structures
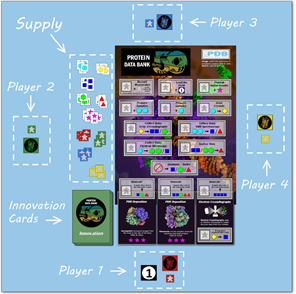
In honor of the PDB’s 50th anniversary, you can now play PDB50: The Game , which illustrates the steps needed to deposit a structure into the PDB, such as:
- Preparing samples: Scientists must isolate the molecule and ensure the sample is not contaminated with other molecules.
- Growing crystals: If using a crystallography technique to determine a structure, scientists must grow crystals of the biological molecules, and this process can be very challenging, time consuming, and sometimes impossible.
- Collecting data: A variety of techniques are available for collecting the data needed to solve a structure.
- X-ray crystallography: the most common technique used, but limited by the ability to crystallize the biological molecule
- Nuclear magnetic resonance spectroscopy: a technique useful for studying biological molecules in liquid so that crystallization isn’t required, but only suitable for small molecules
- Cryo-electron microscopy: a more recently developed technique where samples are flash frozen in solution, requiring no crystallization
- Analyzing data: The data from these techniques don’t give the clean, beautiful images that you see in the PDB; instead, scientists must use computers to process and analyze the data to determine the structure.
- Innovation: In the early days of the PDB, X-ray crystallography was the only method for determining structures. Scientists have since developed, and continue to develop, new methods including:
- X-ray free electron laser, which produces rapid X-ray pulses useful in collecting data from large numbers of microscopic crystals that are too small for standard X-ray crystallography
- Integrative/hybrid methods that can answer complex structural questions by combining computational and multiple experimental techniques
- Electron crystallography, which uses an electron microscope to collect data from tiny crystals
- Automation that allows scientists to maximize their sample preparation efficiency
- PDB deposition! Once the structure of a protein or other biological molecule is determined, scientists can submit it to the PDB for other scientists to access and study.
Success of the PDB
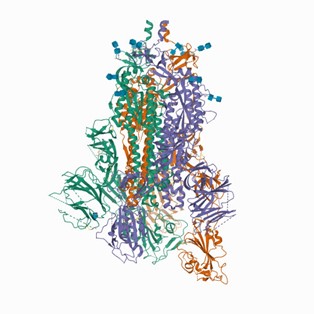
Even though determining the structure of a biological molecule is difficult and time consuming, knowing these structures is essential for understanding their roles in human health and disease. Proof of the PDB’s impacts is evident in the many Nobel Prizes that have been awarded to scientists for their work on or with molecular structures. The structures in the PDB are also very useful for discovering new medications. Knowing the specific structure of the targeted protein allows scientists to fine-tune the drug’s chemistry to optimize its effectiveness. For example, PDB resources have played a vital role in providing researchers with information about proteins on the surface of SARS-CoV-2, the virus responsible for the COVID-19 pandemic. This knowledge helped researchers determine the most promising candidates for drug and vaccine development.
You can also celebrate the PDB’s 50th anniversary the next time you play your favorite card game by downloading these structural biology playing cards, and learn more about the PDB’s history and future from this 50th anniversary video. For more examples of protein structures, check out our interactive protein alphabet activity!
The PDB is supported in part by NIGMS grant R01GM133198.

This post is a great supplement to Pathways: The Imaging Issue.
Researchers use many imaging techniques in determining protein (and other macromolecular) structures. Some of these techniques are described in Pathways, and others are introduced in this post.
Learn more in our Educator’s Corner.


