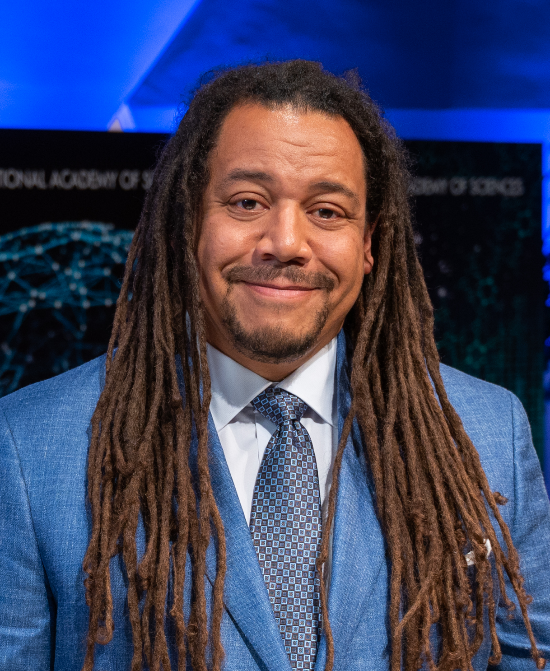
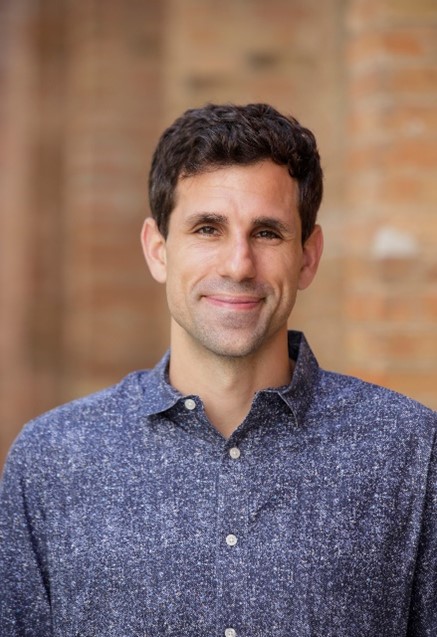
“I found a passion for both biology and chemistry in high school and thought, Well, that must mean I’m a biochemist! Luckily my naïve thought was correct. I am a biochemist,” says Bil Clemons,
Ph.D. He’s a professor of biochemistry at the California Institute of Technology (Caltech) in Pasadena, where he’s been teaching and running a lab for nearly 20 years.
A Path to Research
Dr. Clemons doesn’t remember a time when he wasn’t interested in science or curious about the world. “I think, fundamentally, that’s what being a scientist is: being curious about how the world works,” he says. As a child, he’d open seed pods to see the insides or take toys apart to see how their tiny motors worked. He couldn’t always figure out how to put the toys back together, though, which led to his parents warning him not to ruin his siblings’ new toys on Christmas morning.
Continue reading “Bil Clemons: Following Scientific Curiosity”