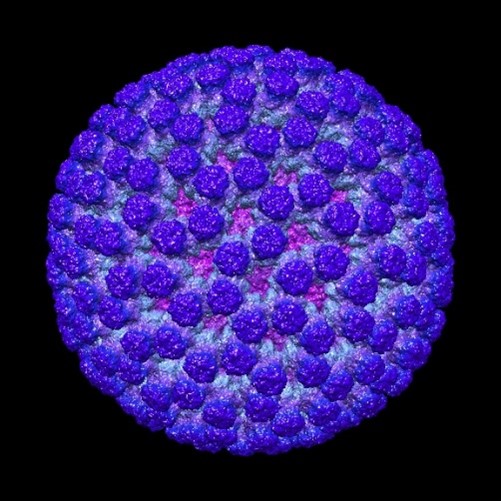
Have you ever noticed a skin care product advertised as “microbiome friendly” and wondered what that meant? The microbiome is the collection of all the microbes—including bacteria, viruses, and fungi—that live in a specific environment, such as on the skin or in the digestive tract.
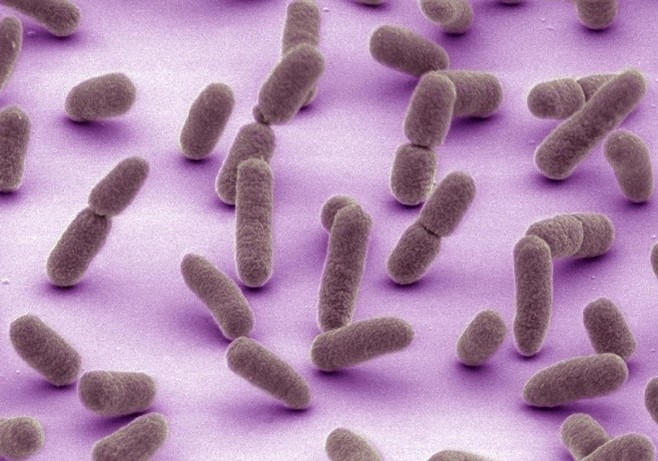
Credit: Mark Ellisman and Thomas Deerinck, National Center for Microscopy and Imaging Research, University of California, San Diego.
It’s a common misconception that all microbes are harmful—in truth, much of the human microbiome is made up of microbes that form beneficial symbiotic relationships with us. Microbiome-friendly skin care products don’t have antimicrobial properties that could harm the beneficial bacteria that live on our skin.
Your Microbiome and You
Continue reading “What Is the Microbiome?”