Visualizations can give scientists unprecedented views of complex biological processes. Here’s a look at two new ones that shed light on how HIV enters host cells.
Animation of HIV’s Entry Into Host Cells
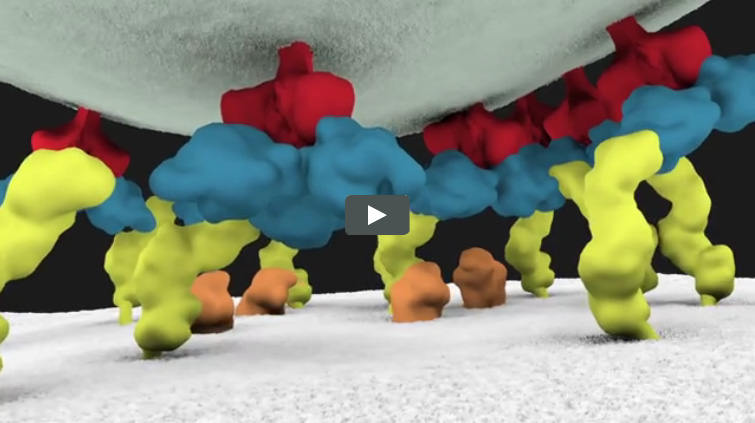

We previously introduced you to Janet Iwasa, a molecular animator who’s visualized complex biological processes such as cells ingesting materials and proteins being transported across a cell membrane. She has now released several animations from her current project of visualizing HIV’s life cycle ![]() . The one featured here shows the virus’ entry into a human immune cell.
. The one featured here shows the virus’ entry into a human immune cell.
“Janet’s animations add great value by helping us consider how complex interactions between viruses and their host cells actually occur in time and space,” says Wes Sundquist, who directs the Center for the Structural Biology of Cellular Host Elements in Egress, Trafficking, and Assembly of HIV ![]() at the University of Utah. “By showing us how different steps in viral replication must be linked together, the animations suggest hypotheses that hadn’t yet occurred to us.”
at the University of Utah. “By showing us how different steps in viral replication must be linked together, the animations suggest hypotheses that hadn’t yet occurred to us.”
To help other researchers explore complex biological processes, Iwasa and her colleagues recently launched Molecular Flipbook, an open-source software toolkit for making molecular animations.
Model of HIV’s Surface Proteins
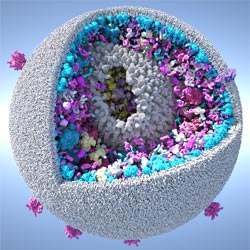
Graham Johnson, a medical illustrator for more than 15 years, created this model of an HIV particle using an open-source software program called cellPACK ![]() that he developed as a graduate student at The Scripps Research Institute (TSRI).
that he developed as a graduate student at The Scripps Research Institute (TSRI).
The cellPACK visualizations reconcile experimental results about the distribution of “spike” proteins on the surface of the immature HIV viral particle. These proteins help the virus enter host cells. The cellPACK models reveal that the arrangement of the proteins is not random, as had been previously suggested. This conclusion, according to the research paper, shows the power of the software to simulate and interpret results from other studies. Plus, the improved understanding of the spike proteins will further elucidate the HIV life cycle and ways to disrupt it.
Getting a closer look at HIV is just one application of the software. In a news release about the software, Art Olson, director of the HIV Interaction and Viral Evolution Center at TSRI and Johnson’s graduate school advisor, says, “We hope to ultimately increase scientists’ ability to target any disease.”

