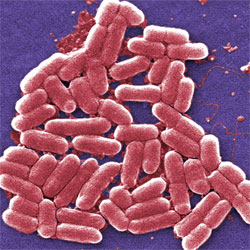
E. coli bacteria help us digest our food, produce vitamin K and have served as a model organism in research for decades. Now, they might one day be harnessed as environmental or medical sensors and long-term data storage devices ![]() .
.
MIT researchers Timothy Lu and Fahim Farzadfard modified the DNA of E. coli cells so that the cells could be deployed to detect a signal (for example, a small molecule, a drug or the presence of light) in their surroundings. To create the modified E. coli, the scientists inserted into the bacteria a custom-designed genetic tool.
When exposed to the specified signal, the tool triggers a series of biochemical processes that work together to introduce a single mutation at a specific site in the E. coli’s DNA. This genetic change serves to record exposure to the signal, and it’s passed on to subsequent generations of bacteria, providing a continued record of exposure to the signal. In essence, the modified bacteria act like a hard drive, storing biochemical memory for long periods of time. The memory can be retrieved by sequencing the bacteria or through a number of other laboratory techniques.
The researchers call the new technology SCRIBE, for Synthetic Cellular Recorders Integrating Biological Events. It’s not only able to detect the presence of a chosen signal, but also the magnitude and duration of exposure.
SCRIBE could be used to indicate chemical contaminants, pollutants or other substances in the environment. Conceivably, it could also serve as a sensor for virtually any molecule in the body—blood proteins, dietary products, immune factors, hormones, disease markers, pharmaceuticals, toxins—to monitor health, detect disease and track the progress of countless medical conditions.
If you’re interested in SCRIBE’s technical details, which include harnessing an enzyme from another type of bacteria, producing a specific RNA-DNA hybrid and generating single-stranded DNA containing the desired mutation, you can read a perspective ![]() on the paper or watch an animation
on the paper or watch an animation ![]() of the process.
of the process.
This work was funded in part by NIH under grants P50GM098792 and DP2OD008435.


Thanks for sharing this nice article and I wish to visit again on your blog. Keep sharing your work.