Cells are the basis of the living world. Our cells make up the tissues and organs of our bodies. Bacteria are also cells, living sometimes alone and sometimes in groups called biofilms. We think of cells mostly as staying in one spot, quietly doing their work. But in many situations, cells move, often very quickly. For example, when you get a cut, infection-fighting cells rally to the site, ready to gobble up bacterial intruders. Then, platelet cells along with proteins from blood gather and form a clot to stop any bleeding. And finally, skin cells surrounding the wound lay down scaffolding before gliding across the cut to close the wound.
This remarkable organization and timing is evident right from the start. Cells migrate within the embryo as it develops so that body tissues and organs end up in the right places. Harmful cells use movement as well, as when cells move and spread (metastasize) from an original cancer tumor to other parts of the body. Learning how and why cells move could give scientists new ways to guide those cells or turn off or slow down the movement when needed.
Glowing Breadcrumbs
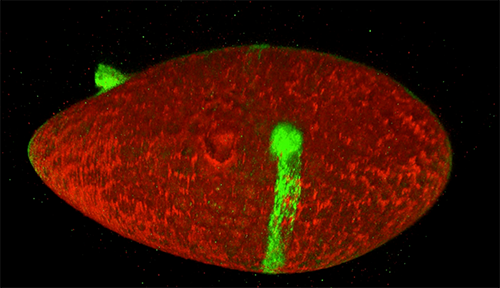
Scientists studying how humans and animals form, from a single cell at conception to a complex body at birth, are particularly interested in how and when cells move. They use research organisms like the fruit fly, Drosophila, to watch movements by small populations of cells. Still, watching cells migrate inside a living fly is challenging because the tissue is too dense to see individual cell movement. But moving those cells to a dish in the lab might cause them to behave differently than they do inside the fly. To solve this problem, NIGMS-funded researcher David Bilder and colleagues at the University of California, Berkeley, came up with a way to alter fly cells so they could track how the cells behave without removing them from the fly. They engineered the cells to lay down a glowing track of proteins behind them as they moved, leaving a traceable path through the fly’s tissue. The technique, called M-TRAIL (matrix-labeling technique for real-time and inferred location), allows the researchers to see where a cell travels and how long it takes to get there.
Bilder and his team first used M-TRAIL in flies to confirm the results of past studies of Drosophila ovaries in the lab using other imaging techniques. In addition, they found that M-TRAIL could be used to study a variety of cell types. The new technique also could allow a cell’s movement to be tracked over a longer period than other imaging techniques, which become toxic to cells in just a few hours. This is important, because cells often migrate for days to reach their final destinations.
Eventually, Bilder believes M-TRAIL could be used to trace the first movements of metastatic cancer cells in humans from a primary tumor out to a blood vessel and surrounding tissues. Learning how cancer cells take these first steps could uncover new ways to stop their migration.
Feet, Fans or Waves? Changing Cellular Modes of Motion
Besides tracking where cells go when they migrate, scientists need to know what cues set them in motion and what controls their movement. NIGMS grantee Peter Devreotes and colleagues at the Johns Hopkins University School of Medicine, Baltimore, MD, are studying a network of connected signaling proteins inside cells. Their findings show that this network is key to controlling cell movement.
Once it’s turned on, this protein network starts up the cell’s movement machinery, including its cellular skeleton, called the cytoskeleton. In response to signals from the network, the cell begins to change shape, an early step in cell migration.
The movement network can start up at random, but it also responds to cues from outside the cell. Devreotes and his team tested this by putting the amoeba cells in a dish of swirling water. Amoeba cells need to be able to respond to liquid moving around them as well as to other signals coming from the surrounding environment. The researchers found that the network proteins within the cells responded to the swirling water in the same way that they respond to other environmental cues. Devreotes thinks the network gives the cell a sophisticated way of interpreting and adjusting to real-world situations.
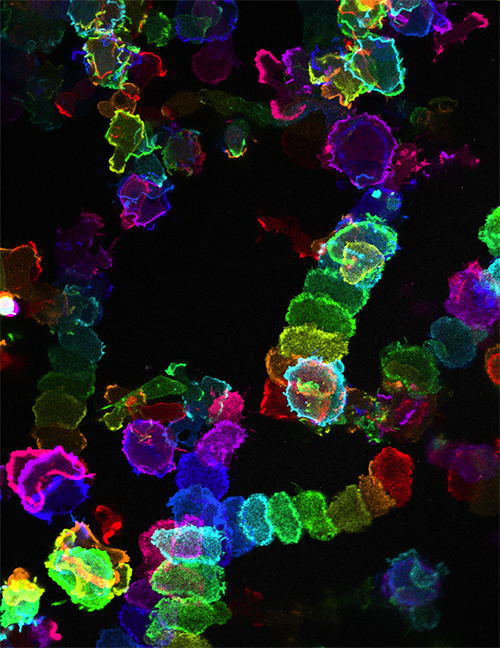
Once on the move, different cells use different techniques to propel themselves forward. Some, like amoeba or our own white blood cells, reach out with bumps called pseudopods, cellular “feet” that pull the cell forward. Others create a large fan-shaped bulge all along the leading edge of the cell that drives it along, and still others make waves from their centers that ripple through the cell, pulling them forward.
Scientists had thought that each kind of cell used only one of these modes to move. Instead, when Devreotes and his team changed the proteins in the movement network inside cells they found that amoebas altered their movement behavior, switching from pseudopods to fans to rippling waves and back again. So, the network Devreotes studies might be both powerful and universal. It may be similar across many kinds of cells from different species, coordinating signals from outside and determining what mechanism the cells will use to move. Further defining this network could help reveal ways to tweak cell movement when it goes awry.
Saving Resources: Every Cell for Itself
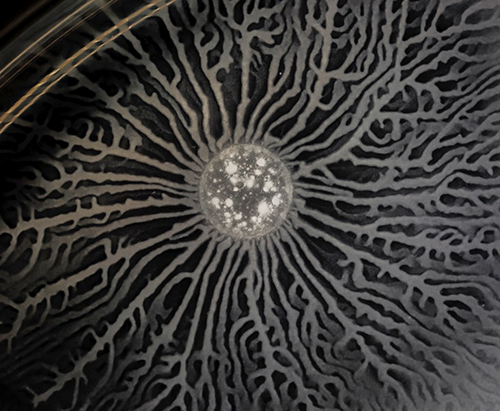
Understanding how individual cells move tells only part of the story. Cells often move as a group, and when they do, they can do complex jobs that you might not expect. For example, instead of using an every-cell-for-itself approach, bacteria can coordinate their behavior to move the entire colony in an orderly way. Scott Chimileski, an NIGMS-funded research fellow and imaging specialist in the laboratory of Roberto Kolter at Harvard University School of Medicine, uses time-lapse photography to observe how bacteria do this.
Moving as a group has benefits for bacteria, but it is also expensive. For example, making and secreting surfactant, a substance that reduces the tension of the surface and makes it easier to slide across, takes a lot of energy for each cell. So, bacteria want to make sure that none of their competitors can use the surfactant they make for a free ride. Bacteria have evolved many methods for telling their close relatives from their enemies. They even release antibiotics that they are immune to but that will kill bacterial cells not related to them. Kolter and his team believe that the weapons bacteria use to compete successfully also make them into close-knit colonies and promote cooperation. Learning about the balance between cooperation and competition in bacterial colonies, Chimileski said, will help clarify what happens in places like our own gut microbiomes, which, we are learning, are highly complex microbial environments very important to our health.
Dr. Bilder’s research is supported by NIH Grants 5R01GM068675-13 and 5R01GM111111-04. Dr. Devreotes’s research is supported by NIH Grant 5R35GM118177-02, and Dr. Kolter’s research is supported by NIH Grant R01GM058213-20.

