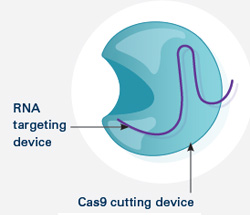
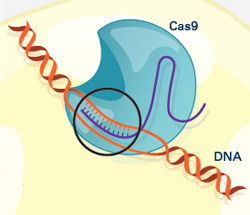
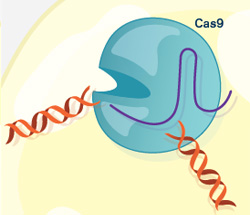
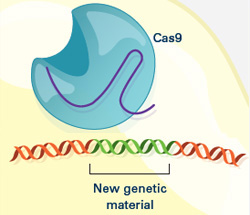
The CRISPR gene-editing tool was recognized today by Science magazine as its “breakthrough of the year.” We support a number of researchers working in this exciting area and have featured it on this blog. To learn more about this exceptionally promising new method, see below for our illustrated explanation of the CRISPR system and its possible applications.
Tag: Cool Tools/Techniques
Cracking a Ubiquitous Code
We asked the heads of our scientific divisions to tell us about some of the big questions in fundamental biomedical science that researchers are investigating with NIGMS support. This article is the third in an occasional series that explores these questions and explains how pursuing the answers could advance understanding of important biological processes.
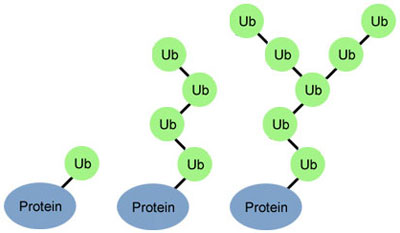
Researchers are on a quest to crack a code made by ubiquitin, a small protein that plays a big role in coordinating cellular function. By attaching to other proteins, ubiquitin determines what those proteins should do next.
Just as zip codes direct letters to specific towns, the ubiquitin code might direct one protein to help with DNA repair, another to assist in cell division, and a third to transport molecules into and out of cells.
Continue reading “Cracking a Ubiquitous Code”Cool Image: Tracing Proteins in Action
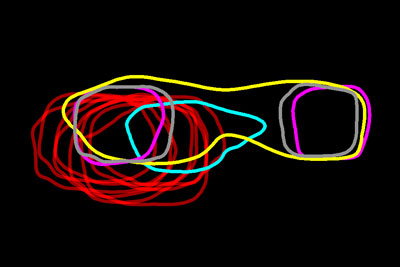

Looking like necklaces stacked on a dresser, these bright, amorphous loops show the outlines of yeast proteins that make up the spindle pole, a cellular component found in organisms as diverse as yeast and humans. Each cell starts with a single spindle pole, which must somehow duplicate to form the pair that works together to pull matching chromosomes apart during cell division. Scientists don’t completely understand how this duplication occurs, but they do know that errors in spindle pole copying can lead to a number of health conditions, including cancer.
Continue reading “Cool Image: Tracing Proteins in Action”Cool Image: DNA Origami
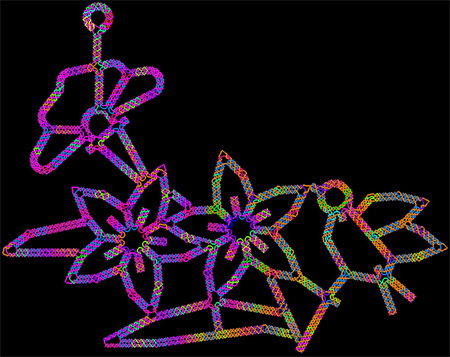
This image of flowers visited by a bird is made of DNA, the molecule that provides the genetic instructions for making living organisms. It shows the latest capability of a technique called DNA origami to precisely twist and fold DNA into complex arrangements, which might find future use in biomedical applications.
Continue reading “Cool Image: DNA Origami”Bacterial ‘Fight Clubs’ and the Search for New Medicines
Bacteria hold a vast reservoir of compounds with therapeutic potential. They use these compounds, known as secondary metabolites, to protect themselves against their enemies. We use them in many antibiotics, anti-inflammatories and other treatments.
Scientists interested in developing new medicines have no shortage of places to look for secondary metabolites. There are an estimated 120,000 to 150,000 bacterial species on Earth. Each species is capable of producing hundreds of secondary metabolites, but often only under specific ecological conditions. The challenge for researchers is figuring out how to coax the bacteria to produce these compounds.
Now, Brian Bachmann and John McLean of Vanderbilt University and their teams have shown that by creating “fight clubs” where bacteria compete with one another, they can trigger the bacteria to make a wide diversity of molecules, including secondary metabolites. Continue reading “Bacterial ‘Fight Clubs’ and the Search for New Medicines”
Mapping Our Skin’s Microbes and Molecules
Last month, we shared some facts about the microbes that inhabit us. Here’s another: From head to toe, our skin bacteria coexist with chemicals in hygiene products, fibers from clothes and proteins shed by dead or dying skin cells.
These images highlight the complex composition of our body’s largest organ. They show the association between microbial diversity (top images) and skin chemistry (middle images). The different colors note the abundance of a certain bacterium or molecule—red is high, and blue is low. The skin maps remind NIH Director Francis Collins of a 60’s rock album cover.
Continue reading “Mapping Our Skin’s Microbes and Molecules”Simulating the Potential Spread of Measles
- Go to http://fred.publichealth.
pitt.edu/measles
- Select “Get Started”
- Pick a state and city
- Play both simulations
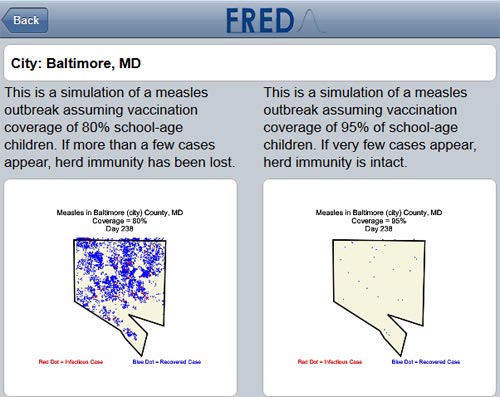
To help the public better understand how measles can spread, a team of infectious disease computer modelers at the University of Pittsburgh has launched a free, mobile-friendly tool that lets users simulate measles outbreaks in cities across the country.
The tool is part of the Pitt team’s Framework for Reconstructing Epidemiological Dynamics, or FRED, that it previously developed to simulate flu epidemics. FRED is based on anonymized U.S. census data that captures demographic and geographic distributions of different communities. It also incorporates details about the simulated disease, such as how contagious it is.
Continue reading “Simulating the Potential Spread of Measles”
Remotely and Noninvasively Controlling Genes and Cells in Living Animals

One of the items on biomedical researchers’ “to-do” list is devising noninvasive ways to control the activity of specific genes or cells in order to study what those genes or cells do and, ultimately, to treat a range of human diseases and disorders.
A team of scientists recently reported progress on a new, noninvasive system that could remotely and rapidly control biological targets in living animals. The system can be activated remotely using either low-frequency radio waves or a magnetic field. Similar radio wave technology operates automatic garage-door openers and remote control car keys and is used in medicine to control electronic pacemakers noninvasively. Magnetic fields are used to activate sensors in burglar alarm systems and to turn your laptop to hibernate mode when the cover is closed. Continue reading “Remotely and Noninvasively Controlling Genes and Cells in Living Animals”
Meet Maureen L. Mulvihill

Credit: Actuated Medical, Inc.
Fields: Materials science, logistics
Works at: Actuated Medical, Inc., a small company that develops medical devices
Second job (volunteer): Bellefonte YMCA Swim Team Parent Boost Club Treasurer
Best skill: Listening to people
Last thing she does every night: Reads to her 7- and 10-year-old children until “one of us falls asleep”
If you’re a fan of the reality TV show Shark Tank, you tune in to watch aspiring entrepreneurs present their ideas and try to get one of the investors to help develop and market the products. Afterward, you might start to think about what you could invent.
Maureen L. Mulvihill has never watched the show, but she lives it every day. She is co-founder, president and CEO of Actuated Medical, Inc. (AMI), a Pennsylvania-based company that develops specialized medical devices. The devices include a system for unclogging feeding tubes, motors that assist MRI-related procedures and needles that gently draw blood.
AMI’s products rely on the same motion-control technologies that allow a quartz watch to keep time, a microphone to project sound and even a telescope to focus on a distant object in a sky. In general, the devices are portable, affordable and unobtrusive, making them appealing to doctors and patients.
Mulvihill, who’s trained in an area of engineering called materials science, says, “I’m really focused on how to translate technologies into ways that help people.” Continue reading “Meet Maureen L. Mulvihill”
Untangling a Trending Topic
It’s not every day that we log into Facebook and Twitter to see conversations about denaturing proteins and the possibility of reducing biotechnology costs, but that changed last week when a story about “unboiling” eggs became a trending topic.
Since NIGMS partially funded the research advance ![]() that led to the media scramble, we asked our scientific expert Jean Chin to tell us more about it.
that led to the media scramble, we asked our scientific expert Jean Chin to tell us more about it.
What’s the advance?
Gregory Weiss of the University of California, Irvine, and his collaborators have designed a device that basically unties proteins that have been tangled together. Continue reading “Untangling a Trending Topic”