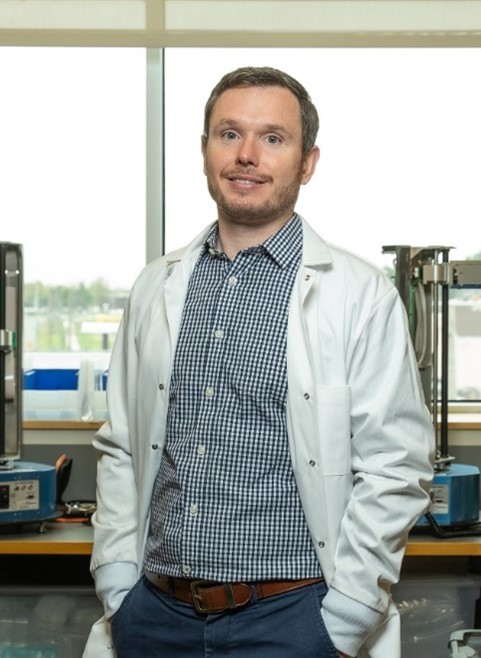
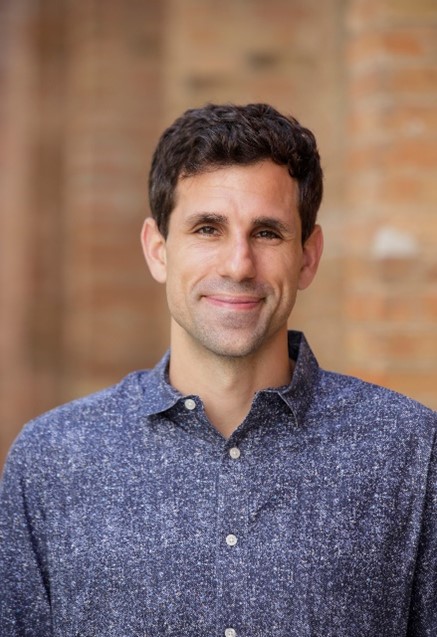
Adam Gormley, Ph.D., describes himself as a creative and adventurous person—albeit, not creative in the traditional sense. “Science allows me to be creative; to me, it’s a form of art. I love being outdoors, going on sailing trips, and spending time adventuring with my family. Research is the same—it’s an adventure. My creative and adventurous sides have combined into a real love for science,” he says. Dr. Gormley currently channels his passion for science into his position as an assistant professor of biomedical engineering at Rutgers University in Piscataway, New Jersey.
Learning How the World Works
Both of Dr. Gormley’s parents worked in science and medicine—his mother as a medical doctor and his father as a physician-scientist—and they instilled in him a curiosity for how the world worked. When he was young, Dr. Gormley and his parents would tinker with cars or boats and fix broken household items together, all the while talking about the individual parts and how they functioned as a whole. “I always had that technical, hands-on side of me,” he says.
Continue reading “Using Robots and Artificial Intelligence to Search for New Medicines”