Ravi Iyengar
Fields: Systems pharmacology and systems biology
Works at: Mount Sinai School of Medicine, New York, NY
Favorite sports team: Yankees
Favorite subject in high school: Math
Recently read book: The Signal and the Noise by Nate Silver
Credit: Pedro Martinez, Systems Biology Center New York
Ravi Iyengar, a professor at Mount Sinai School of Medicine, stood in an empty lecture hall, primed to tell thousands of students about systems biology, a holistic approach to studying fundamental life processes. To prepare for this moment, he had spent 4 months reading hundreds of scientific papers and distilling the research into understandable nuggets. But that day, his only student was a videographer.
Together, they recorded 15 different lectures about systems biology—many related to Iyengar’s own research—that thousands of people would stream or download as part of a MOOC, or massive open online course.
Trained in biochemistry, Iyengar built his research career around studying molecules and developing a list of all the parts that help nerve, kidney and skin cells to function. As he obtained more information, he realized he needed to know how all the components worked together. To achieve this comprehensive understanding, Iyengar turned to computational techniques and mathematical analyses—cornerstones of systems biology.
For more than a decade, he has been using and developing systems biology approaches to explore a range of biomedical questions, from very basic to translational ones with immediate relevance to human health.
Iyengar’s Findings
In his earlier work, Iyengar used mathematical analyses to show that molecules within cells connect with one another to form switches that produce cellular memory. This may allow, for instance, an immune cell to remember a foreign object and secrete an antibody. In recent work, he and his team developed a mathematical model showing that the shape of a cell influences the flow of information across the membrane, possibly contributing to disease states and offering a way to study and identify them under the microscope. In another study, they analyzed a database of drug side effects to find combinations of medications that produce fewer adverse reactions and then created a cell biology interaction network that explains why a certain drug pair had this beneficial outcome. The approach could point to other combinations of FDA-approved drugs that reduce serious side effects and thereby guide clinical practice.
“Systems biology is a powerful way to explore important biological and medical questions, and it’s relevant to many fields of science,” said Iyengar. But he added that the majority of educational institutions, including liberal arts and community colleges, don’t have systems biology courses. So, Iyengar teamed with colleagues to create a series of MOOCs.
The first course, offered last summer and taught by Iyengar, presented all the facets of systems biology. The syllabus included lessons on genomics and bioinformatics, fields that have contributed to systems biology; gathering and integrating data; and the use of modeling in drug development.
“My goal was for the students to get the general gestalt of systems biology,” explained Iyengar, who directs an NIH-funded center focused on the systems-level study of medicine and therapeutics.
In total, more than 12,000 participants watched at least one video lecture, 3,000 submitted one or more of the weekly quizzes and 1,800 took a mid-term or final exam. The online discussions forum included nearly 400 topics with about 5,000 posts. The students, most enrolled in a graduate program or working full-time, had some training in the biological, biomedical, computer and information sciences.
“The stats tell me that many people are in fields adjacent to systems biology and don’t have access to more traditional systems biology courses,” concluded Iyengar. “Through the MOOC, we can reach them in a substantial way.”
The second course, which covers network analysis, wrapped up in early December, and the third course, which covers dynamical modeling methods, began in January. Iyengar plans to offer the intro course again in late March.
Learn more:
MOOC Systems Biology Courses 



 View the
View the 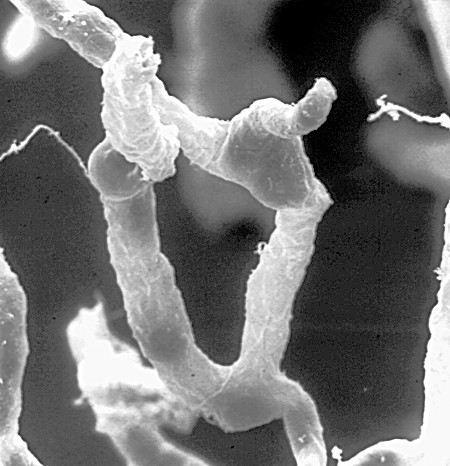 A network of capillaries supplies brain cells with nutrients. Tight seals in their walls keep blood toxins—and many beneficial drugs—out of the brain. Credit: Dan Ferber, PLOS Biol 2007 Jun; (5)6:E169.
A network of capillaries supplies brain cells with nutrients. Tight seals in their walls keep blood toxins—and many beneficial drugs—out of the brain. Credit: Dan Ferber, PLOS Biol 2007 Jun; (5)6:E169.  Credit: University of Nebraska, Lincoln.
Credit: University of Nebraska, Lincoln.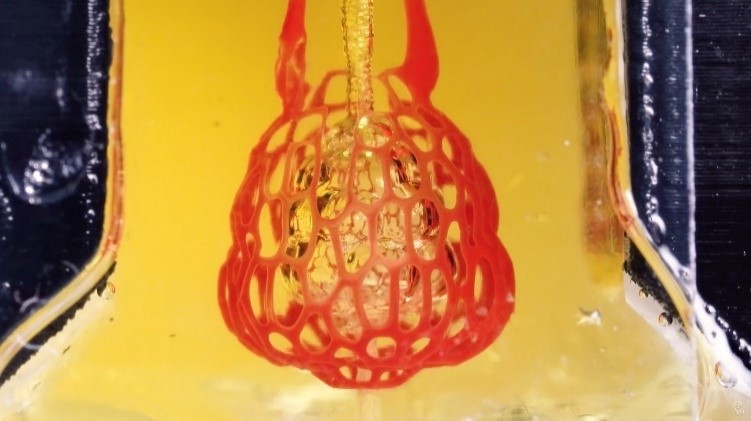 A bioprint of the small air sac in the lungs with red blood cells moving through a vessel network supplying oxygen to living cells. Credit: Rice University.
A bioprint of the small air sac in the lungs with red blood cells moving through a vessel network supplying oxygen to living cells. Credit: Rice University.


