As Halloween approaches, we turned up some spectral images from our gallery. The collection below highlights some spooky-sounding—but really important—biological topics that researchers are actively investigating to spur advances in medicine.


Biomedical Beat Blog – National Institute of General Medical Sciences
Follow the process of discovery
As Halloween approaches, we turned up some spectral images from our gallery. The collection below highlights some spooky-sounding—but really important—biological topics that researchers are actively investigating to spur advances in medicine.

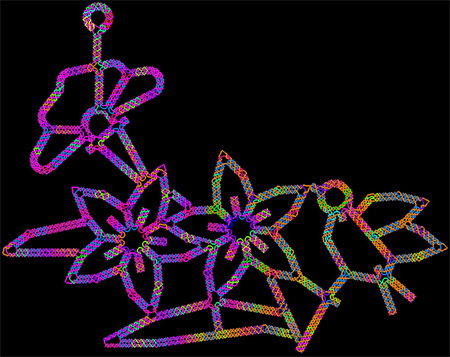
This image of flowers visited by a bird is made of DNA, the molecule that provides the genetic instructions for making living organisms. It shows the latest capability of a technique called DNA origami to precisely twist and fold DNA into complex arrangements, which might find future use in biomedical applications.
Continue reading “Cool Image: DNA Origami”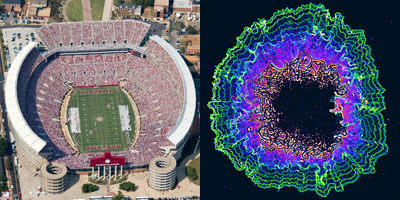
What do these images of football fans and bacterial cells have in common? By following simple rules, each individual allows the group to accomplish tasks none of them could do alone—a stadium wave that ripples through the crowd or a cell colony that rebounds after antibiotic treatment.
These collective behaviors are just a few examples of what scientists call emergent phenomena. While the reasons for the emergence of such behavior in groups of birds, fish, ants and other creatures is well understood, they’ve been less clear in bacteria. Two independent research teams have now identified some of the rules bacterial cells follow to enable the colony to persist. Continue reading “The Simple Rules Bacteria Follow to Survive”
Last month, we shared some facts about the microbes that inhabit us. Here’s another: From head to toe, our skin bacteria coexist with chemicals in hygiene products, fibers from clothes and proteins shed by dead or dying skin cells.
These images highlight the complex composition of our body’s largest organ. They show the association between microbial diversity (top images) and skin chemistry (middle images). The different colors note the abundance of a certain bacterium or molecule—red is high, and blue is low. The skin maps remind NIH Director Francis Collins of a 60’s rock album cover.
Continue reading “Mapping Our Skin’s Microbes and Molecules”Visualizations can give scientists unprecedented views of complex biological processes. Here’s a look at two new ones that shed light on how HIV enters host cells.
Animation of HIV’s Entry Into Host Cells
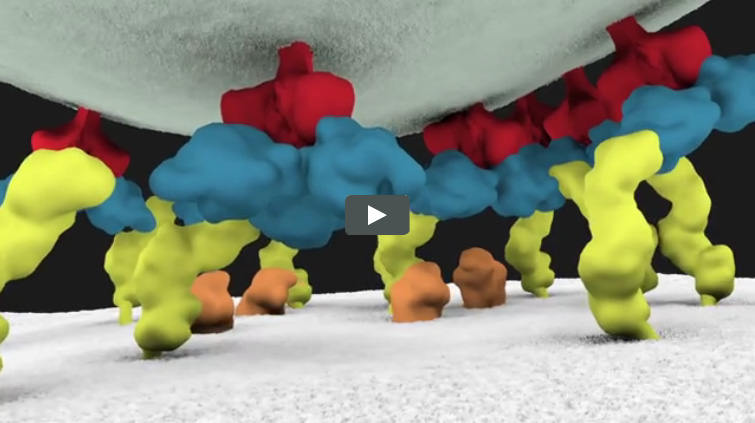

We previously introduced you to Janet Iwasa, a molecular animator who’s visualized complex biological processes such as cells ingesting materials and proteins being transported across a cell membrane. She has now released several animations from her current project of visualizing HIV’s life cycle ![]() . The one featured here shows the virus’ entry into a human immune cell.
. The one featured here shows the virus’ entry into a human immune cell.
“Janet’s animations add great value by helping us consider how complex interactions between viruses and their host cells actually occur in time and space,” says Wes Sundquist, who directs the Center for the Structural Biology of Cellular Host Elements in Egress, Trafficking, and Assembly of HIV ![]() at the University of Utah. “By showing us how different steps in viral replication must be linked together, the animations suggest hypotheses that hadn’t yet occurred to us.”
at the University of Utah. “By showing us how different steps in viral replication must be linked together, the animations suggest hypotheses that hadn’t yet occurred to us.”
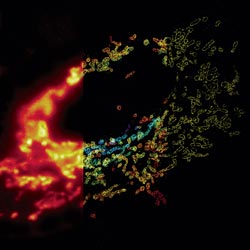
Much as a photographer brings distant objects into focus with a telephoto lens, scientists can now see previously indistinct cellular components as small as a few billionths of a meter (nanometers). By overcoming some of the limitations of conventional optical microscopy, a set of techniques known as super-resolution fluorescence microscopy has changed once-blurry images of the nanoworld into well-resolved portraits of cellular architecture, with details never seen before in biology. Reflecting its importance, super-resolution microscopy was recognized with the 2014 Nobel Prize in chemistry.
Using the new techniques, scientists can observe processes in living cells across space and time and study the movements, interactions and roles of individual molecules. For instance, they can identify and track the proteins that allow a virus to invade a cell or those that enable tumor cells to migrate to distant parts of the body in metastatic cancer. The ability to analyze individual molecules, rather than collections of molecules, allows scientists to answer longstanding questions about cellular mechanisms and behavior, such as how cells move along a surface or how certain proteins interact with DNA to regulate gene activity. Continue reading “Field Focus: Bringing Biology Into Sharper View with New Microscopy Techniques”
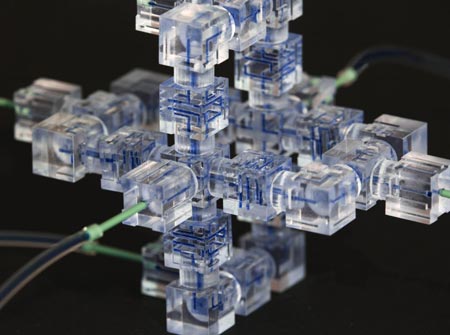

Like snapping Lego blocks together to build a fanciful space station, scientists have developed a new way to assemble a microfluidics system, a sophisticated laboratory tool for manipulating small volumes of fluids.
Microfluidics systems are used by scientists to perform tasks as diverse as DNA analysis, microbe detection and disease diagnosis. Traditionally, they have been slow and expensive to produce, as each individual “lab on a chip” had to be built from scratch in a special facility.
Now, researchers including Noah Malmstadt of the University of Southern California have harnessed 3-D printing technology to create a faster, cheaper, easier-to-use system ![]() . The team first identified the smallest functional pieces of a microfluidics system. Each of these pieces performs one simple task like detecting the size of fluid droplets or mixing two fluids together. After 3-D printing individual components, the team showed that they could be snapped together by hand into a working system in a matter of hours. The individual pieces can be pulled apart and re-assembled as needed before use in an actual experiment, which was impossible with the traditional microfluidics systems.
. The team first identified the smallest functional pieces of a microfluidics system. Each of these pieces performs one simple task like detecting the size of fluid droplets or mixing two fluids together. After 3-D printing individual components, the team showed that they could be snapped together by hand into a working system in a matter of hours. The individual pieces can be pulled apart and re-assembled as needed before use in an actual experiment, which was impossible with the traditional microfluidics systems.
The researchers have created eight block-like components so far. They hope to start an online community where scientists will share designs for additional components in an open-source database, helping to speed further development of the technology.
This work was funded in part by NIH under grant R01GM093279.
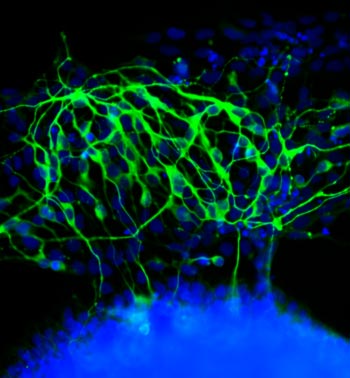
If you think this image looks like the fluorescent outline of a brain, you’re on the right track. The green threads show neurons that have just formed from unspecialized cells called stem cells.
Researchers led by Laura Kiessling ![]() of the University of Wisconsin-Madison directed the stem cells to become neurons by changing the quality of the surface on which they grew. In experiments testing different gels used to grow stem cells in the lab, the scientists found that the stiffness of those gels influenced cell fate decisions.
of the University of Wisconsin-Madison directed the stem cells to become neurons by changing the quality of the surface on which they grew. In experiments testing different gels used to grow stem cells in the lab, the scientists found that the stiffness of those gels influenced cell fate decisions.
When grown on a soft gel with a brain tissue-like surface, the stem cells began to transform into neurons. This happened without the addition of any of the proteins normally used to coax stem cells to specialize into different types of cells.
A better understanding of how stem cell fate is influenced by the mechanical properties of a surface could help researchers who are trying to harness the blank slate cells for tissue regeneration or other therapeutic uses.
This work also was funded by NIH’s National Institute of Biomedical Imaging and Bioengineering; National Heart, Lung, and Blood Institute; and National Institute of Neurological Disorders and Stroke.
Learn more:
University of Wisconsin-Madison News Release ![]()
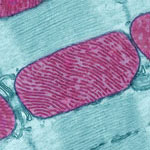
Meet mitochondria: cellular compartments, or organelles, that are best known as the powerhouses that convert energy from the food we eat into energy that runs a range of biological processes.
As you can see in this close-up of mitochondria from a rat’s heart muscle cell, the organelles have an inner membrane that folds in many places (and that appears here as striations). This folding vastly increases the surface area for energy production. Nearly all our cells have mitochondria, but cells with higher energy demands have more. For instance, a skin cell has just a few hundred, while the cell pictured here has about 5,000.
Scientists are discovering there’s more to mitochondria than meets the eye, especially when it comes to understanding and treating disease.
Read more about mitochondria in this Inside Life Science article.
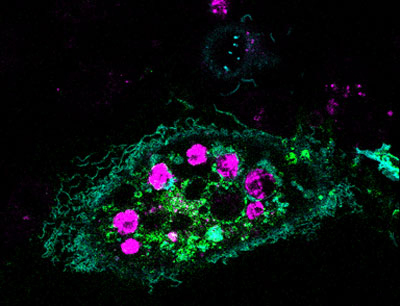
In this image, a healthy cell (green) has engulfed a number of dying cells (purple) just as a predator might ingest wounded or dying prey. A team of researchers is hoping to use this same strategy at the cellular level to help treat infection, neurodegenerative diseases or cancer.
Our bodies routinely use a process called phagocytosis to rid themselves of unhealthy cells. Takanari Inoue at Johns Hopkins University and collaborators at the University of Tokyo set out to investigate the molecular underpinnings of phagocytosis. Their goal was to endow laboratory-grown human cells with phagocytotic skills, namely the ability to recognize, swallow and digest dying cells. The scientists tried to do this by inserting into the cells two molecules known to play a role in phagocytosis.
The engineered cells accomplished the first two tasks—they recognized and surrounded dying cells. But they didn’t digest what they’d engulfed. Now the researchers are looking for a molecular trigger to get the engineered cells to complete this last task.
Eventually, the scientific team aims to build artificial cells that are programmed to target and destroy abnormal cells, such as those ravaged by bacteria, cancer or other diseases.
Learn more:
Johns Hopkins News Release ![]()
How Cells Take Out the Trash Article from Inside Life Science
Cellular Suicide: An Essential Part of Life Article from Inside Life Science